Difference between revisions of "Slicer3:Module:QueryAtlas"
| Line 47: | Line 47: | ||
* '''Annotation & Display Options panel:''' | * '''Annotation & Display Options panel:''' | ||
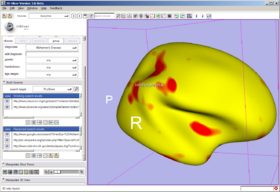
| + | In the Annotation & Display Options, select among termsets to be used to generate the interactive annotations in the 3D Viewer (local terms, BIRNLex Strings, NeuroNames Strings, IBVD terms, and UMLS Concept Names). Anatomical regions that are mapped from the local termset to any of the others will be annotated on mouse-over; if no translation of the local term is available, no annotation will be displayed. The Scene Visibility menu can be used to turn on and off annotations in the 3D viewer, and to toggle model visibility, which can be useful for viewing slice planes and label map or statistical overlays that a model may occlude. | ||
| + | |||
* '''Ontology Mapping panel:''' | * '''Ontology Mapping panel:''' | ||
* '''Search Terms panel:''' | * '''Search Terms panel:''' | ||
Revision as of 21:13, 13 October 2007
Home < Slicer3:Module:QueryAtlasQueryAtlas
Slides for Oct07 BIRN meeting
General Information
Module Type & Category
Type: Interactive
Category: N/A
Authors & Collaborators
Steve Pieper, Greg Brown, David Kennedy, Maryanne Martone, Jyl Boline, Burak Ozyurt,Wendy Plesniak, Michael Halle, Anna Tang, and Florin Talos
Module Description
The QueryAtlas module can be used to:
- Load volumetric analyses from XCEDE compliant or XNAT databases;
- Visualize volumes, models and parametric map overlays.
- Annotate models and labelmaps with atlas-based anatomy labels
- Map those local anatomy names to symantically compatible terms used in the larger biomedical informatics (neuroinformatics) community
- Formulate queries in the context of particular study results (fMRI activations or group statistics, for example)
- View, manage and save collections of query results
Usage
Example data
Download FIPS/FreeSurfer Xcede catalog example data: QueryAtlasExampleData.zip .
Download Qdec (XNAT) example data: TEST5.qdec .
Quick Tour of Features and Use
- Scene Setup panel:
In the FIPS/FreeSurfer tab, load FIPS/FreeSurfer morphological and functional analyses from Xcede 2.0 catalog files, arrange data in the foreground, background and label layer of the Slice Viewers through the interface, and set up annotations on the datasets. Complete datasets should include a high resolution anatomical scan (brain.mgz), a label map generated by FreeSurfer's automatic parcellation (aparc+aseg.mgz), the pial surface models (lh.pial and rh.pial) an example functional dataset used in the FIPS analysis (example_func.nii) and resulting brain activation statistics of interest (zstat.nii, zfstat.nii), and the matrix FreeSurfer uses to register the example functional and anatomical image (anat2efx.register.dat).
In the Qdec tab, load a Qdec population statistics analysis results file (file.qdec) extracted from an XNAT archive, select from among the scalar overlays representing the statistics in the study, and set up annotations on the dataset.
- Annotation & Display Options panel:
In the Annotation & Display Options, select among termsets to be used to generate the interactive annotations in the 3D Viewer (local terms, BIRNLex Strings, NeuroNames Strings, IBVD terms, and UMLS Concept Names). Anatomical regions that are mapped from the local termset to any of the others will be annotated on mouse-over; if no translation of the local term is available, no annotation will be displayed. The Scene Visibility menu can be used to turn on and off annotations in the 3D viewer, and to toggle model visibility, which can be useful for viewing slice planes and label map or statistical overlays that a model may occlude.
- Ontology Mapping panel:
- Search Terms panel:
- Build Queries panel:
Development
Dependencies
Other modules or packages that are required for this module's use.
Known bugs
Follow this link to the Slicer3 bug tracker:
http://na-mic.org/Mantis/main_page.php
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing:
http://na-mic.org/Mantis/main_page.php
Source code & documentation
Customize following links for your module:
http://www.na-mic.org/ViewVC/index.cgi/
Links to documentation generated by doxygen:
http://www.na-mic.org/Slicer/Documentation/Slicer3/html/
More Information
Acknowledgement
This research was supported by Grant 5 MOI RR 000827 to the FIRST BIRN and Grant 1 U24 RR021992 to the FBIRN Biomedical Informatics Research Network (BIRN, http://www.nbirn.net), that is funded by the National Center for Research Resources (NCRR) at the National Institutes of Health (NIH). This work was also supported by NA-MIC, NAC, NCIGT. NeuroNames ontology and URI resources are provided courtesy of BrainInfo, University of Washington.