Difference between revisions of "2011 Summer Project Week Integrate BRAINSCut into Slicer3"
From NAMIC Wiki
m |
|||
| (2 intermediate revisions by the same user not shown) | |||
| Line 14: | Line 14: | ||
==Key Investigators== | ==Key Investigators== | ||
| − | * University of Iowa: Eunyoung(Regina) Kim | + | * University of Iowa: Eunyoung(Regina) Kim, Hans Johnson, Mark Scully |
| Line 22: | Line 22: | ||
<h3>Objective</h3> | <h3>Objective</h3> | ||
| − | # | + | #Integration of BRAINSCut into Slicer3/Slicer4 |
| − | # | + | #Integration of GMI feature images into BRAINSCut |
| − | # | + | #Test BRAINSCut |
</div> | </div> | ||
| Line 31: | Line 31: | ||
<h3>Approach, Plan</h3> | <h3>Approach, Plan</h3> | ||
| − | # | + | # Check dependencies of BRAINSCut and integrate into Slicer CLM. |
| − | + | #Process test data set to verify | |
| − | + | # Submit caudate results to the CAUSE07 Challenge 'http://www.cause07.org/' | |
</div> | </div> | ||
</div> | </div> | ||
| Line 39: | Line 39: | ||
<h3>Progress</h3> | <h3>Progress</h3> | ||
| − | + | # BRAINSCut Incorporating with Slicer3 DONE | |
| + | # Generating BRAINSCut model for subcortical structures: caudate, putamen, thalamus, and hippocampus. DONE | ||
| + | # Verifying generated model by comparing manual traces. DONE | ||
| + | # Process 230 data to generate hippocampus. DONE | ||
| Line 47: | Line 50: | ||
#Powell, S., Magnotta, V., Johnson, H., and . . . , V. J., “Registration and machine learning-based automated segmentation of subcortical and cerebellar brain . . . ,” NeuroImage (Jan 2008) | #Powell, S., Magnotta, V., Johnson, H., and . . . , V. J., “Registration and machine learning-based automated segmentation of subcortical and cerebellar brain . . . ,” NeuroImage (Jan 2008) | ||
| − | #Eun Young Kim, Hans Johnson, "Multi-structure segmentation of multi-modal brain images using artificial neural networks", SPIE Proceedings Paper(Mar 2010) | + | #Eun Young Kim, Hans Johnson, "Multi-structure segmentation of multi-modal brain images using artificial neural networks", SPIE Proceedings Paper (Mar 2010) |
==Download Page== | ==Download Page== | ||
Latest revision as of 14:36, 24 June 2011
Home < 2011 Summer Project Week Integrate BRAINSCut into Slicer3Integrate BRAINSCut into Slicer3
Key Investigators
- University of Iowa: Eunyoung(Regina) Kim, Hans Johnson, Mark Scully
Objective
- Integration of BRAINSCut into Slicer3/Slicer4
- Integration of GMI feature images into BRAINSCut
- Test BRAINSCut
Approach, Plan
- Check dependencies of BRAINSCut and integrate into Slicer CLM.
- Process test data set to verify
- Submit caudate results to the CAUSE07 Challenge 'http://www.cause07.org/'
Progress
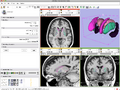
- BRAINSCut Incorporating with Slicer3 DONE
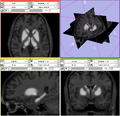
- Generating BRAINSCut model for subcortical structures: caudate, putamen, thalamus, and hippocampus. DONE
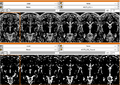
- Verifying generated model by comparing manual traces. DONE
- Process 230 data to generate hippocampus. DONE
References
- Powell, S., Magnotta, V., Johnson, H., and . . . , V. J., “Registration and machine learning-based automated segmentation of subcortical and cerebellar brain . . . ,” NeuroImage (Jan 2008)
- Eun Young Kim, Hans Johnson, "Multi-structure segmentation of multi-modal brain images using artificial neural networks", SPIE Proceedings Paper (Mar 2010)
Download Page
http://www.nitrc.org/projects/brainscut
Delivery Mechanism
This work will be delivered to the NAMIC Kit as a
- NITRIC distribution
- Slicer Module
- Built-in:
- Extension -- commandline:
- Extension -- loadable: