Difference between revisions of "2009 Summer Project Week Project Segmentation of Muscoskeletal Images"
| Line 2: | Line 2: | ||
<gallery widths="400px" perrow="6"> | <gallery widths="400px" perrow="6"> | ||
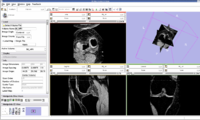
Image:Knee.jpg| Knee MRI Image | Image:Knee.jpg| Knee MRI Image | ||
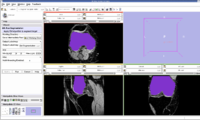
| − | Image: | + | Image:LabelMap Femur.jpg|Filled Femur Label Map. |
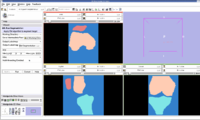
Image:Femur Patella Tibia.jpg|EM Segmented Output. | Image:Femur Patella Tibia.jpg|EM Segmented Output. | ||
</gallery> | </gallery> | ||
Revision as of 17:29, 29 May 2009
Home < 2009 Summer Project Week Project Segmentation of Muscoskeletal Images
Key Investigators
- Stanford: Harish Doddi, Saikat Pal, Scott Delp
- Harvard: Ron Kikinis
- Steve Pieper, Isomics, Inc.
Objective
The aim of this project is to develop an automatic/semi-automatic methodology to convert whole body imaging datasets into three-dimensional models for neuromuscular biomechanics and finite element simulations.
Approach, Plan
We are working on understanding the capabilities of RegisterImage module in Slicer to apply to knee datasets. Currently we are conducting parameter exploration studies to evaluate the sensitivity of registered images to different input parameters associated with the algorithms. We are also developing a module to apply python ICP-based registration algorithms to directly morph a surface model to a target image geometry.
Our goals for the project week are:
- Perform and evaluate results from an extensive
parameter space exploration study of RegisterImages Batchmake module on knee dataset.
- Resolve issues in building Python modules from
slicer source code.
- Demonstrate proof of concept on registering an
existing atlas (.vtk, .stl) to a target image using Python ICP Registration module.
Progress