Difference between revisions of "DBP3:Utah:AutoEndoSeg"
From NAMIC Wiki
| Line 14: | Line 14: | ||
= Initial Experiences = | = Initial Experiences = | ||
| − | * Comparison of | + | * Comparison of GA Tech and Utah-derived segmentations |
(click to enlarge) | (click to enlarge) | ||
Revision as of 08:30, 29 November 2011
Home < DBP3:Utah:AutoEndoSegContents
Notes on the Endocardial Segmentation Algorithm
Algorithm is a multi-atlas segmentation.
- Register (cropped) heart region of patient DEMRI to all of the n atlas DEMRI images and record n minimum metric values (mutual information)
- Transform all n atlas segmentations to patient DEMRI space
- Segmentation is a weighted average of the transformed atlas segmentation, where weights are determined by the mutual information values from step 1
- Pick a threshold as your segmentation
Notes on Yi's Slicer Module
- The exposed parameter is the ratio of pixel samples for mutual information to the total number of pixels in the image.
- We could also expose the threshold parameter in the final step
- We can change or add atlas datasets by copying into the appropriate directory and editing the text file that lists them
- Uses the ITK mutual information code. Does affine registration followed by b-spline
- We will partner with Yi for module development at the Jan 2012 project week
Initial Experiences
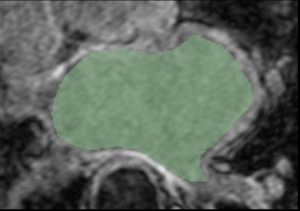
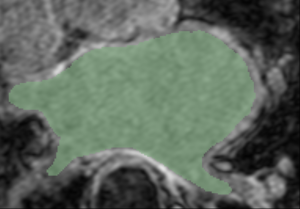
- Comparison of GA Tech and Utah-derived segmentations
(click to enlarge)
| GA Tech Segmentation | Utah Segmentation | Thresholding Difference |
|---|---|---|
| Segmentation of an atlas image by GA Tech, using their endocardial auto-segmentation algorithm. | Segmentation of the same image with the same algorithm by Utah. | User-based variability in thresholding the probability mask output. (Green region - Utah; white region - GA Tech). |