DBP2:UNC:Regional Cortical Thickness Pipeline
From NAMIC Wiki
Revision as of 20:47, 17 October 2008 by Mathieuc (talk | contribs) (→Pipeline description (steps))
Home < DBP2:UNC:Regional Cortical Thickness Pipeline
Back to UNC Cortical Thickness Roadmap
Objective
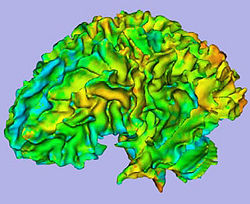
We would like to create an end-to-end application within Slicer3 allowing individual and group analysis of regional cortical thickness.
General information
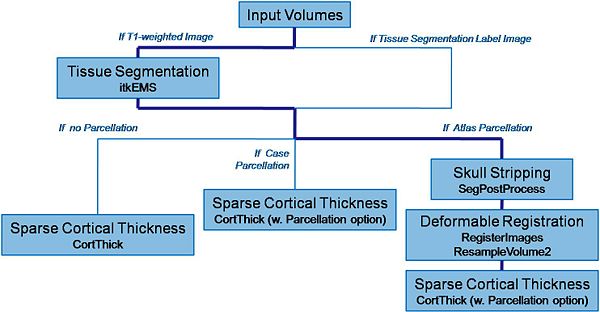
Pipeline description (steps)
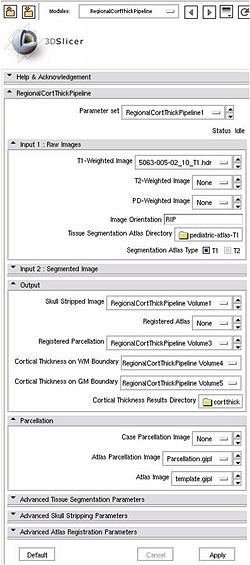
Input: T1-weighted image, T2-weighted image, PD-weighted image
- 1. Tissue segmentation
- Tool: itkEMS (UNC Slicer3 external module)
- 2. Skull stripping
- Tool: SegPostProcess (UNC Slicer3 external module)
- 3. Deformable registration of pediatric regional atlas
- 3.1 Deformable registration of T1-weighted pediatric atlas
- Tool: RegisterImages (Slicer3 module)
- 3.2. Applying transformation to the parcellation map
- Tool: ResampleVolume2 (Slicer3 module)
- 3.1 Deformable registration of T1-weighted pediatric atlas
- 4. Cortical Thickness
- Tool: CortThick (UNC Slicer3 module)
- 1. Tissue segmentation
All the tools used in the current pipeline are Slicer3 modules, some of them being UNC external modules. The user can thus compute a regional cortical thickness analysis on one data, either within Slicer3 or by running the pipeline as command lines.
Usage (Command Line)
Inputs: T1-weighted image, T1-weigthed atlas, regional atlas (parcellation map)
A. Pipeline command line
RegionalCortThickPipeline --T1 Image_T1.gipl --segAtlasDir TissueSegmentationAtlasDirectory/ --atlas Atlas.gipl --atlasParcellation Parcellation.gipl --SaveWM WMCorticalThicknessMap --SaveGM GMCorticalThicknessMap
B. Step by step command line
- 1. Tissue segmentation
- Input: EMSparam.xml
- Output: Image_Corrected_EMS.gipl, Label.gipl
- 1. Tissue segmentation
itkEMSCLP --XMLFile EMSparam.xml
- 2. Skull stripping
- Input: Label.gipl, Image_Corrected_EMS.gipl
- Output: Image_Corrected_EMS_Stripped.gipl, BinaryMask.gipl (optional)
- 2. Skull stripping
SegPostProcessCLP Label.gipl Image_Corrected_EMS_Stripped.gipl --skullstripping Image_Corrected_EMS.gipl
- 3. Deformable registration of pediatric regional atlas
- 3.1 Deformable registration of T1-weighted pediatric atlas
- Input: Atlas.gipl, Image_Corrected_EMS_Stripped.gipl
- Output: Atlas_Registered.gipl, Atlas_Registered_Transform.txt
- 3.1 Deformable registration of T1-weighted pediatric atlas
- 3. Deformable registration of pediatric regional atlas
RegisterImages Image_Corrected_EMS_Stripped.gipl Atlas.gipl –resampledImage Atlas_Registered.gipl –saveTransform Atlas_Registered_Transform.txt –registration PipelineBSpline
- 3.2. Applying transformation to the parcellation map
- Input: Parcellation.gipl, Atlas_Registered_Transform.txt, Image_Corrected_EMS_Stripped.gipl
- Output: Parcellation_Registered.gipl
- 3.2. Applying transformation to the parcellation map
ResampleVolume2 Parcellation.gipl Parcellation_Registered.gipl -f Atlas_Registered_Transform.txt -i nn -R Image_Corrected_EMS_Stripped.gipl
- 4. Cortical Thickness
- Input: Parcellation_Registered.gipl, Label.gipl
- Output: CortThick_Result_Dir/, WMCorticalThicknessMap, GMCorticalThicknessMap
- 4. Cortical Thickness
CortThickCLP CortThick_Result_Dir/ --par Parcellation_Registered.gipl --inputSeg Label.gipl --SaveWM WMCorticalThicknessMap --SaveGM GMCorticalThicknessMap
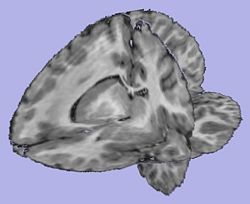
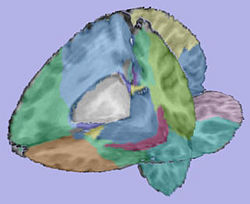
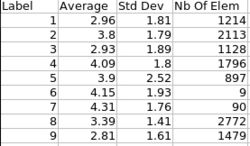
Analysis on a small pediatric dataset
Tests have been computed on a small pediatric dataset, including 2 years old and 4 years old cases
- 2 Autistic cases
- 1 developmental delay
- 1 normal control
In progress
- Workflow for individual analysis (Slicer3 external module using BatchMake)
Future work
- UNC Slicer3 external modules available on NITRIC
- Pediatric atlas (T1-weighted image and parcellation map) available to the community (XNAT?)
- Workflow for group analysis