Projects:ARRA:SlicerEM
Contents
Aim
The EMSegmenter is a state-of-the-art segmentation tool within 3D Slicer. User feedback has reported that clinicians are currently unable to tune the approach to their acquisition protocol, as the user interface is too complex. This proposal addresses this issue by redesigning the user interface, focusing on hiding the complexity of the underlying segmentation algorithm. If successful, this will enable clinicians to automatically segment their own medical scans, even if the corresponding acquisition protocol deviates from the default setting for which the EMSegmenter is optimized.
Research Plan
The EMSegmenter is the result of 15 years of research in medical image segmentation. This has lead to a user interface that exposes a rich set of parameters. These parameters allow the tuning of the EMSegmenter to a wide variety of acquisition sequences. However, tuning these parameters is quite challanging. In addition, Slicer currently does not provide any tools for generating atlases, which are a set of parameters characterizing each structure of interest. We propose to address these issues by creating two user-friendly modules: one for generating the atlases and one for tuning the EMSegmenter to a specific acquisition sequence.
The first module, called Atlas Generator, builds the atlases characterizing each structure of interest. The user simply specifies the training data which can be done via querying XNAT, a database targeted towards medical image analysis. The user also selects the type of information to be extracted from the data. Possible types are the shape, intensity, or relative position of the structures of interest across the training set. Based on this input, the tool automatically generates the atlas.
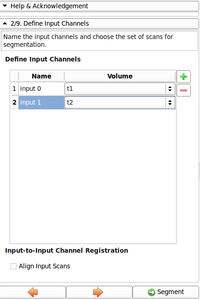
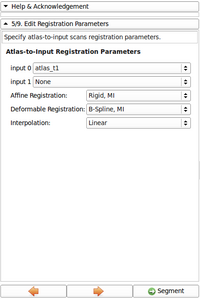
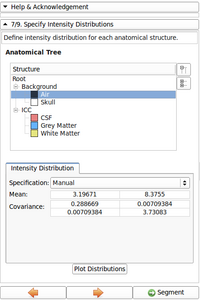
The second module, called EMSegmenter-Simple, consists of a simple work flow that enables users to adjust the EMSegmenter to their specific acquisition sequence. As part of this proposal, we will create a library of templates, which parametrizes the tool to segmentation tasks frequently ancountered by our user community. User simply adopt the tool to their acquisition scenario by first selecting the proper template. In the second step, the user modifies the value of important parameters of the template. We simplify the tuning of these parameters by providing an instant feedback mechanism. The feedback mechanism updates the automatic segmentation according to the change in the parameter setting. This will allow users to to get an intuitive understanding about impact of certain parameters on the algorithm. In addition, each entry field will be accompanied with a help text. We will also create detailed documentation about the user interface and publish a tutorial for each template.
The project is viewed as successful if a properly trained clinician is able to modify the templates to their acquisition sequence within an hour.
Events
- Advanced EMSegmenter Training : How to parametrize the tool
- Where: 1249 Bolyston Street, Boston, MA
- When: 10 am - 1 pm , Feb 23, 2010
Key Personnel
20% Kilian Pohl
95% Dominique Belhachemi
5% Daniel Haehn
Kitware (Project Manager: Jean-Christophe Fillion-Robin)
Documentation
- EMSegmenter documentation for Slicer 3.6
- Developer page
Progress
- 07/02/11
- Installed MacOS and WindowsXP test machines
- It is now possible to use a temporary directory with white spaces (this was a bug on Windows XP)
- Added manual sampling to EMSegmenter module
- 04/02/11
- option intermediate results directory is created for the user
- clarified the usage, helped collaborators with the EMSegmenter
- Windows specific runtime bug fixed, happened whenever we used an iterator two times.
- Included detailed verbose printout in EMSegmentCommandLine
- Added test results to MIDAS server
- Use CMake to compare segmentation results with precomputed segmentation results
- Further cleanup code (Number of registration flags simplified)
- Simplified MRML structure(removed nodes)
- Check if atlas files are properly downloaded to avoid FetchMI checksum checks
- Fixed bug in test EMSegCL_IntermediateResults: "UpdateScene: error getting 0th storage node"
- Updated template creator to new MRML structure (new MRML structure)
- 04/02/11
- Fixed the updated functionality for EMSegmenter
- also transferred fix to Extension wizard (was very quick)
- added new repository for 3.6.3 for EMSegment tasks
- fix for the new webserver index files and therefore do not need the index.html
- committed all code to 3.6.3 Branch and Trunk
- tested on Mac and Linux
- Atlas Creator Updates:
- Created diagram of Atlas Creator workflow
- Support for Normalization of output Atlases done
- Support for Selecting the Output Cast done
- Re-Designed API
- Focus on Cluster proposal
- Focus on external invocation (command line or 3rd party module in 3D Slicer)
- Introduced Configuration Concept
- Documentation, Diagram and Priority List updated at Projects:ARRA:SlicerEM:AtlasCreator
- Fixed the updated functionality for EMSegmenter
- 28/01/11
- Fixed a bug in the test 'intermediateResultsDirectory'. The test doesn't use the results file which is specified in the mrml scene. Instead it is using a test specific results file(specified in the CMake template).
- Fixed a Slicer4 bug : "ENH: Call the GeneralGauss(...) functions instead of the FastGauss(...) functions - this avoids type punning" Modules/EMSegment/Graph/vtkImageCurveRegion.cxx [15896]
- Fixed a Slicer4 bug : "BUG: Use a normal gauss function to avoid type-punning" QTModules/EMSegment/Widgets/vtkPlotGaussian.cxx
- Worked on the CT-Hand-Bone task
- Worked on a EMSegmenter specific skull stripper
- Atlas Creator Updates:
- Research and Proposal of running the Atlas Creator in a cluster environment
- Several GUI Fixes
- Support Dynamic Registration (in the works)
- Support for using existing Registrations by loading deformation fields (in the works)
- Support for Normalization of output Atlases (in the works)
- Support for Selecting the Output Cast (in the works)
- Merged Export of Deformation Fields and Export of Transforms
- 21/01/11
- Worked on new CT-Hand-Bone task
- Maintaining work on existing tasks
- Added new features to the Atlas Creator
- New wiki page Projects:ARRA:SlicerEM:AtlasCreator
- 14/01/11
- Created a test for the MRI-Human-Brain-Full-Parcellation task
- Worked on failing tests if compiling Slicer3 with Superbuild
- Worked on new CT-Hand-Bone task
- Worked with Wendy on new ideas how to make FetchMI more rubust
- 07/01/11
- Created new Atlas Creator module for Slicer3
- Created new Wiki page for Call for Datasets for the EM Segmenter Use Case Library
- 12/31/10
- Help debugging FetchMI bugs 1027 983
- Updated MRI-Human-Brain-Parcellation
- 12/24/10
- Add postprocessing to segmentation, which includes island removal and subparcellation of tissue classes
- 12/17/10
- Added down-sampled image data for testing purposes
- Added two tests for each task
- Added UPenn logo to the EMSegmenter acknowledgment section
- Work on FetchMI bugs
- Updated Wiki page for the task MRI-Human-Brain-Parcellation
- 12/10/10
- Added wrapper for SkullStripperCLI to GenericTask.tcl
- Updated CMTK extension, using now Torsten Rohlfing's repository: https://nitrc.org/svn/cmtk/tags/Slicer3
- Updated Advanced Tutorial based on Ron's comments
- Added functionality to threshold target image(s), negative values will be set to 0 so that the EMSegmenter can be used on those images
- 12/03/10
- Create a new Wiki page for the EMSegmenter Task Library [1]
- Updated task files and documentation
- MRI-Human-Brain
- MRI-Human-Brain-Parcellation
- Non-Human-Primate
- Fixed a apache server issue to update tasks in Slicer 3.6.2
- 11/26/10
- Fixed bug: when creating new task that registration was performed regardless if it was selected or not
- Worked on test cases for 'Non Human Primate' pipeline
- 11/19/10
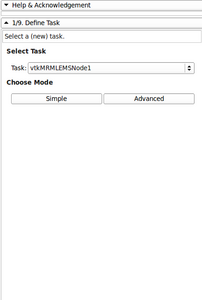
- Created a new EMSegmenter Tutorial (simple mode) - (for quick results, no need to change parameters, this will be useful to clinicians)
- Created a new EMSegmenter Tutorial (advanced mode) - for experienced user of the EMSegmenter
- Created the new 'MRI Human Brain Parcellation' pipeline and added to 3D Slicer 3.6.2 (collaborators: Padmapriya Srinivasan and Sylvain Bouix, PNL BWH)
- 11/12/10
- Restructuring EMSegmenter's Tasks folder to support multiple pipelines
- Start working on a 'MRI Human Brain Parcellation' pipeline (collaborators: Padmapriya Srinivasan and Sylvain Bouix, PNL BWH)
- Start working on a 'Non Human Primate' pipeline (collaborators: Andriy Fedorov, BWH)
- Slicer 3.6.2-RC2 maintenance
- 11/05/10
- software testing for Slicer 3.6.2 release
- Added new test for option --intermediateResultsDirectory
- Fixed bug 1023: MRML import doesn't read copied mrml scenes reliable
- Fixed bug 1025: Slicer crashes when BRAINSFit returns error
- 10/29/10
- Pop up error message when performing segmentation
- Add CMTK in Generic.tcl
- BUG: use .mat extension for transformation files" GenericTask.tcl
- Bug fixing with Slicer Base Code developers
- 10/23/10
- Bug fixes for Slicer 3.6.2 release
- Memory leak fixed
- comment-out unused code in MRI-Human-Brain.tcl
- 10/15/10
- Added function for automatically generating Template File
- Added function for downloading task files from web
- Added CMTK to extension module
- One can assign now tcl file when creating a new task
- Added class name to all tabs
- Tested command line
- Created virtual machines to submit tests reports to the dashboard
- Created Popup window when an error message appears in pre-processing and segmentation
- EMSegmenter works now with different image data types (short, float,...)
- Removed several bugs in mrml structure
- 10/8/10
- Added the --tmp_dir option to the start up allowing one to specify slicers temporary directory
- Fixed the leaks in the nightly tests
- 10/1/10
- Removed command line nodes from mrml scene after the command line is executed
- Added functionality to call N4ITKBiasFieldCorrectionCLI from EMSegmentCommandLine
- Added label - in addition to the tree - to show the selected anatomical tree node
- Patched KWWidgets to specify configuration directory (like /tmp/.3D\ Slicer\ Version\ 3.6.2-betarc)
- Working on parallel execution of multiple instances of the EMSegmentCommandLine
- 9/24/10
- Added check to make sure that the user cannot delete the root node
- Added a warning to avoid ambiguous input channel names
- Added check for emtpy 'Task Name'
- Added a label to show the selected node during specifying the priors
- Alpha value was not correctly transfered from node to vtkImageEMLocalSegmenter
- 9/17/10
- 969 fixed: missing quick timeout if connection to web resource is blocked (closed, see commit 14981+14982)
- 954 fixed:EMSegmentCommandLine throws errors if started from a current working directory different than ./Slicer3-build
- 948+958 fixed:EMSegmentCommandLine segfaults in cleaning up mrmlScene step
- 959 fixed: EMSegmenter ignores option --disableMultithreading
- Added progress dialogs for pre-processing and segmentation
- Test GUI on Linux, Windows, and Apple for Slicer 3.6.2 release
- 9/10/10
- Investigating a critical segfault bug 948+958
- Reviewing use of the extension manager for our needs
- Resolved bugs 970, 959, 944, 943
- 9/3/10
- Updated Image gallery
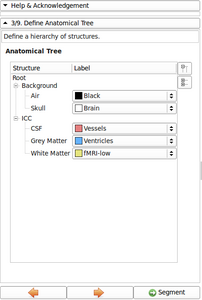
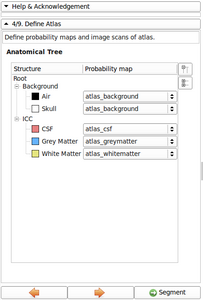
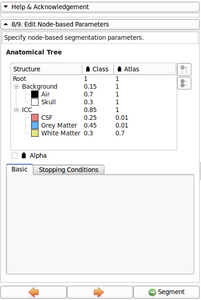
- Refine Anatomical tree widget (Probability map integrated, Icon associated with structure name, ...)
- Now possible to design step directly from QtDesigner
- Discussed integration of PNL kMeans pipeline into Slicer 3.6
- Discussed integration of CNL workflow as a EMSegmenter pipeline into Slicer 3.6
- Discussed differences between the SBIA cluster and the BWH cluster,
- Conclusion: SGE seems to be easier to handle than Lava
- Working on bug 959: EMSegmenter ignores option --disableMultithreading
- 8/27/10
- Integrated Workflow manager with QtModule
- Review Workflow manager API
- Fix bug related to PythonQt wrapping
- Now possible to update atlas class weight, atlas weight and alpha value directly from the Anatomical tree
- Updated EMSegmenter related Wiki pages
- Testing EMSegmenter command line executable in a Sun Grid Engine (SGE) environment (branch 3.6 + trunk)
- working on related bugs 954, 955, 948
- changing some stderr output to stdout
- Updating EMSegmenter tutorial for Slicer 3.6
- 8/20/10
- Ported anatomical tree widget and list of input channel - The anatomical tree is generic enough and re-used.
- Added Image gallery
- 8/13/10
- Meet with Kilian - review panels and widgets and talked about which features should be improved, changed or omitted
- 8/6/10
- Now possible to enable wrapping of QtModule (Gui, UItools) in Slicer through CTK/PythonQt
- Discuss how TCL script interacted with the KWWidgets workflow manager
- 7/30/10
- The port of the Workflow manager is initiated
- Created Qt static panel skeleton corresponding to the step of MRI human brain workflow
- Ability to script task in python is something important. Re-prioritize the task.
- Discussed how dynamic panel could be ported.
- 7/23/10
- Reviewed milestone and list of widgets to port
- Created initial directory structure
- Initiated the conversion of Graph/Algorithm/MRML code to separate shared module
- The port of existing TCL function related to processing is low priority
- 7/16/10
- First weekly meeting between Kitware and UPenn. Meeting will always be held after the Annotation/QT tcon.
- 7/9/10
- Remove bugs
- Set up contract with Kitware to port GUI to QT
- 7/2/10
- Hired Dominique Belhachemi
- Transfered grant from IBM to UPenn
- Wrote Progress Report
- 6/25/10
- Addressed all EMSegmenter bug in Mentis
- 6/18/10
- Revising task for MRI brain segmentation
- Started search for new hire
- Make EMSegmenter work with Slicer Superbuild
- 6/11/10
- Updated EMSegmenter documentation to reflect changes in Slicer 3.6 interface
- Fixed major bugs so that Slicer 3 does not crash when creating a new task
- 6/04/10
- Imbedded BRAINSFit registration in the pipeline
- 5/28/10
- Changed Simple version of EMSegmenter to work with new preprocessing structure
- 5/21/10
- Fixed class overview window
- 5/14/10
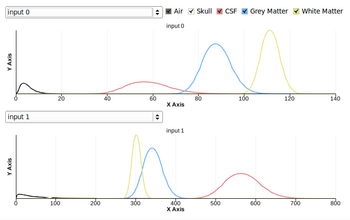
- Fixed Intensity graph distribution display
- 5/07/10
- Fixed EMSegmenter command line module
- 4/30/10
- Debugged EMSegmenter for Slicer 3.6 release
- 4/23/10
- Added segmentation button at each Step of the Wizard
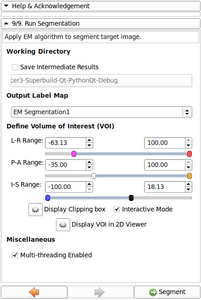
- Integration of 3D/2D Bounding Box selection
- 4/16/10
- Updated Slicer 3.6 Documentation
- Modified workflow of Slicer 3.6
- 4/9/10
- Created simple interface for preprocessing
- 4/2/10
- Incorporated Bug fixes from Slicer 3.4 into trunk
- Removed bugs so that selecting tasks from a list properly works
- Rolled EMSegmenter back to Slicer version 3.4
- EMSegmenter Research environment now compiles under windows
- 3/26/10 Help users of EMSegment to adopt the tools to their needs
- 3/19/10 Manuscript was accepted by CVPR 2010 for poster presentation
- 3/12/10 Start organizing tutorials for EMSegmenter
- 3/05/10 Extended first step in EMSegmenter to include default task list
- 2/26/10 Removed bug so that api interface returns the same results as GUI version in Slicer2.6 . Trained members of CNL, BWH in segmenting lesions from MR brain images. Introduced CNL team to post processing tools for lesion segmentation.
- 2/19/10 Working with Andrey Fedorov begin_of_the_skype_highlighting end_of_the_skype_highlighting begin_of_the_skype_highlighting end_of_the_skype_highlighting begin_of_the_skype_highlighting end_of_the_skype_highlighting begin_of_the_skype_highlighting end_of_the_skype_highlighting begin_of_the_skype_highlighting end_of_the_skype_highlighting, BWH, to customize the EMSegmenter to non-human primate brain images
- 2/12/10 Presented Slicer to IBM Health Care Division
- 2/05/10 Transferring Responsibilities for the EMSegmenter in Slicer 3.5 from MIT to IBM
- 1/29/10 Migrated EMSegmenter Research Environment from Slicer2 to Slicer3
- 1/22/10 Created slides for NAC Site visit
- 1/15/10 Started to resolve bugs that requires collaboration between Kitware, BWH, and IBM team
- 1/8/10 Trained members from BWH and Virginia Tech on using EMSegmenter on Non-Human primate data
- 1/1/10 Addressed all major bugs listed in Mantis for Slicer Version 3.4
- 12/25/09 Setup Slicer environment for new hire
- 12/18/09 Hired Yong Zhang to help with maintenance of EMSegmenter
- 12/11/09 Learned about programming user interfaces in Qt
- 12/04/09 Interviewed applicant for Post Doc position to work on this project
- 11/29/09 Removed bugs related to updated CMake version
- 11/20/09 Organize bugs related to EMSegmenter
- 11/13/09 Tcon featuring the EMSegmenter - participants Andriy Fedorov, Sylvain Jaume, Stuart Wallace, Kilian Pohl - Result of discussion
- Sylvain will fix EMSEgmenter bugs Slicer 3.5
- Kilian will fix any EMSegmenter bugs in Slicer 3.4 and earlier
- Andriy is developing a segmentation pipeline for the Wake Forest Data
- Stuart will do testing of the EMSegmenter module
- 11/06/09 Meet with Jean-Christophe Fillion-Robin from Kitware to discuss integration of Qt in 3D Slicer
- 10/30/09 Organized Monthly TCon between EMSegmenter developers
- 10/23/09 Organized onsite interview , got in contact with Steve Pieper to discuss next steps, installed Slicer3
- 10/17/09 Started interviewing postdoc as well as solving several HR issues for hiring personal
Image gallery
- EM Segmenter - Port to Qt
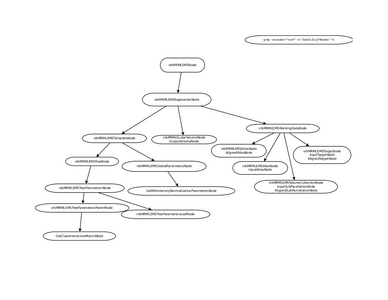
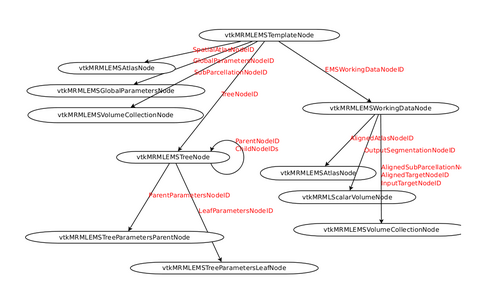
MRML Structure
- EM Segmenter - MRML Structure