Projects:RegistrationLibrary:RegLib C20
From NAMIC Wiki
Home < Projects:RegistrationLibrary:RegLib C20
v3.6.3
Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
Contents
v3.6.3  Slicer Registration Library Case #20: Intra-subject whole-body PET-CT
Slicer Registration Library Case #20: Intra-subject whole-body PET-CT
Input
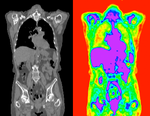
|
|
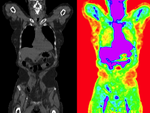
|
| fixed image/target | moving image |
Modules
- Slicer 3.6.3 recommended modules: BrainsFit,
Objective / Background
Change assessment.
Keywords
PET-CT, whole-body, change assessment
Input Data
- reference/fixed : baseline CT: 0.98 x 0.98 x 5 mm , 512 x 512 x 149; PET: 4.1 x 4.1 x 5 mm , 168 x 168 x 149
- moving: CT: 0.98 x 0.98 x 5 mm , 512 x 512 x 149; PET: 4.1 x 4.1 x 5 mm , 168 x 168 x 149
Registration Results
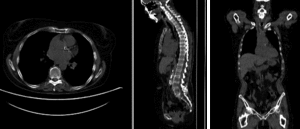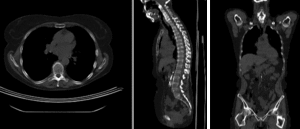 rigid
rigid
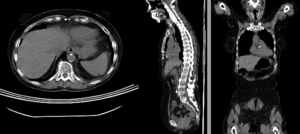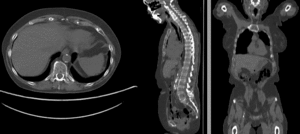 affine alignment
affine alignment
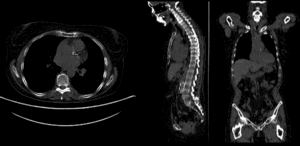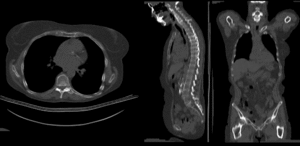 BSpline registration of full volumes. 9 x 9 x 5 grid
BSpline registration of full volumes. 9 x 9 x 5 grid
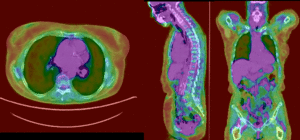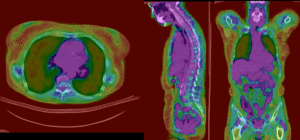 PET overlay ; BSpline registration of full volumes. 9 x 9 x 5 grid
PET overlay ; BSpline registration of full volumes. 9 x 9 x 5 grid
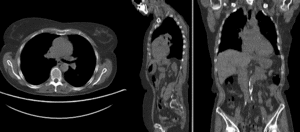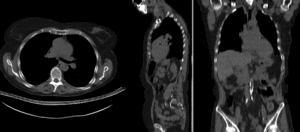 BSpline registration of full volumes. 11 x 11 x 7 grid
BSpline registration of full volumes. 11 x 11 x 7 grid
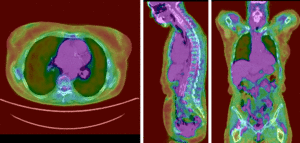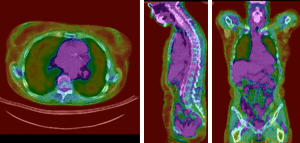 PET overlay ; BSpline registration of full volumes. 11 x 11 x 7 grid
PET overlay ; BSpline registration of full volumes. 11 x 11 x 7 grid
Download
- Data
Procedures
- Phase 1: rigid alignment
- Go to the BRAINSfit module
- select Presets "Xf1_Rigid" or "Xf2_Affine" or set the parameters as given below:
- fixed image: "CT_1", moving image: "CT_2"
- Initialize with previous transform: select "Off"
- Initialize Transform Mode: check box for use MomentsAlign
- check Include Rigid registration Phase box.
- Output: under Slicer Linear Transform, select new and rename to "Xf1_Rigid" or similar
- Registration Parameters: set "Number of Samples" to 200,000 at least
- leave rest at defaults
- Click Apply
- return to the Data module and drag the CT_2 inside/outside the different registration transforms to compare the alignment
- Phase 2: affine alignment
- Go to the BRAINSfit module
- same as Phase 1 above, except:
- Initialize with previous transform: select "Xf1_Rigid" from phase 1 above
- Initialize Transform Mode: check box for Off
- Output: under Slicer Linear Transform, select new and rename to "Xf2_Affine" or similar
- Phase 3: BSpline alignment
- same as Phase 2 above, except:
- Initialize with previous transform: select "Xf2_Affine" from phase 2 above
- Output: under Slicer BSpline Transform, select new and rename to "Xf3_BSpline" or similar
- Output Image Volume: select new and rename to "CT_2_Xf3" or similar
- Registration Parameters: set "Number of Samples" to 250,000 at least
- Number of Grid Subdivisions: 9,9,5 or 11,11,7
- Click Apply
- return to the Data module and drag the CT_2 inside/outside the different registration transforms to compare the alignment
- to obtain a resampled volume: move CT_2 inside Xform of choice and then right-click on the volume and select Harden Transforms. Save MRI under new name.
- Phase 4: Resample PET
- Go to the ResampleScalarVectorDWIVolume module
- Input Volume: PET_2 ; Reference Volume: PET_1 ; Output Volume: create new, rename to "PET_2_Xf3"
- Transform Node: select "Xf3_BSpline" created in phase 3 above
- Transform Order: check box for "output-to-input"
- Click Apply
Discussion: Registration Challenges
- accuracy is the critical criterion here. We need the registration error (residual misalignment) to be smaller than the change we want to measure/detect. Agreement on what constitutes good alignment can therefore vary greatly.
- because of the large FOV we have strong non-rigid deformations from differences in patient/limb positions etc.
- images are large volumes (>100 MB total)
- 2 images pairs have to be aligned, i.e. the calculated transform must be applied to the second (PET) image.
Discussion: Key Strategies
- to calculate the transform, we use the images with the most accurate geometric representation and the smallest expected change, i.e. we align the follow-up CT to the baseline CT and then apply the transforms to the PET image.
- because of the non-rigid differences due to posture and breathing we will need to apply a 2-step registration with an affine alignment followed by a BSpline.