Projects:ShapeBasedSegmentationAndRegistration
Contents
Description
Standard image based segmentation approaches perform poorly when there is little or no contrast along boundaries of different regions. In such cases segmentation is mostly performed manually using prior knowledge of the shape and relative location of the underlying structures combined with partially discernible boundaries. We present an automated approach guided by covariant shape deformations of neighboring structures, which is an additional source of prior knowledge. Captured by a shape atlas, these deformations are transformed into a statistical model using the logistic function. The mapping between atlas and image space, structure boundaries, anatomical labels, and image inhomogeneities are estimated simultaneously within an Expectation-Maximization formulation of the Maximum A posteriori Probability (MAP) estimation problem.
Parameter Estimation Model For Joint Registration and Segmentation
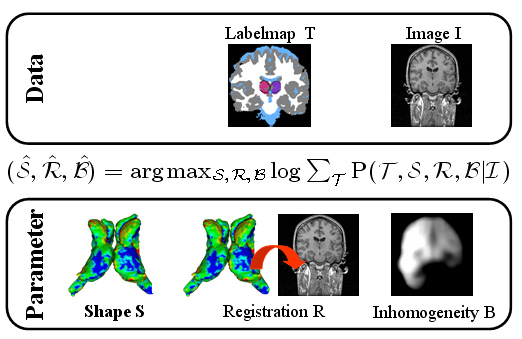
| The Bayesian framework models the relationship between the observed data I (input image), the hidden data T (label map), and the parameter space (S,R,B) with shape model S, registration parameters R, and image inhomogeneities B. The optimal solution with respect to (S,R,B) is defined by the MAP estimate of the framework. We iteratively determine the solution of the estimation problem using an Expectation Maximization (EM) implementation. The E-Step calculates the ‘weights’ for each structure at voxel x. The M-Step updates the approximation of (S,B,R). |
Shape Atlas
|
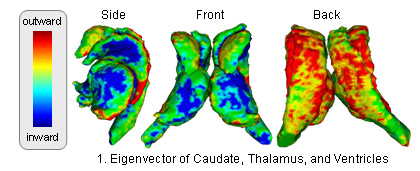
The atlas captures shapes by the signed distance map representation and covariation of the shapes across structures through Principle Component Analysis (PCA). The atlas is constructed through the following method (see [1] for further detail: )
|
Registration Model
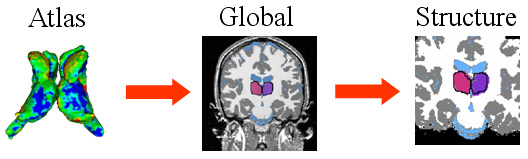
| The hierarchical registration framework represents the correspondence between the coordinate system of the atlas, which defines the shape model, and the MR image. The structure-independent parameters capture the correspondence between atlas and image space. The structure dependent parameters are the residual structure-specific deformations not adequately explained by the structure-independent registration parameters. |
Image Inhomogeneity Model
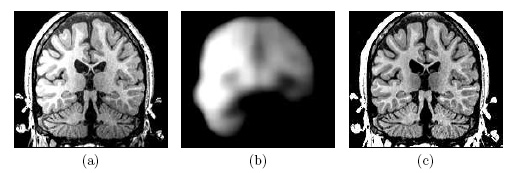
| Image (a) shows an MR image corrupted by image inhomogeneities, noise, partial voluming and other image artifacts. The image inhomogeneities of (a) are shown in (b). Unlike noise, image inhomogeneities are characterized by a slowly varying values within the brain. Image (c) is as image inhomogeneity corrected MR image of (a). In our method, inhomogeneities are modeled as a Gaussian distribution over the image space. As Wells et al. [2] shows the inhomogeneity can then be approximated by the product between a low pass filter represented as a matrix and the weighted residual between estimated and observed MR image. |
Results
Example Segmentation
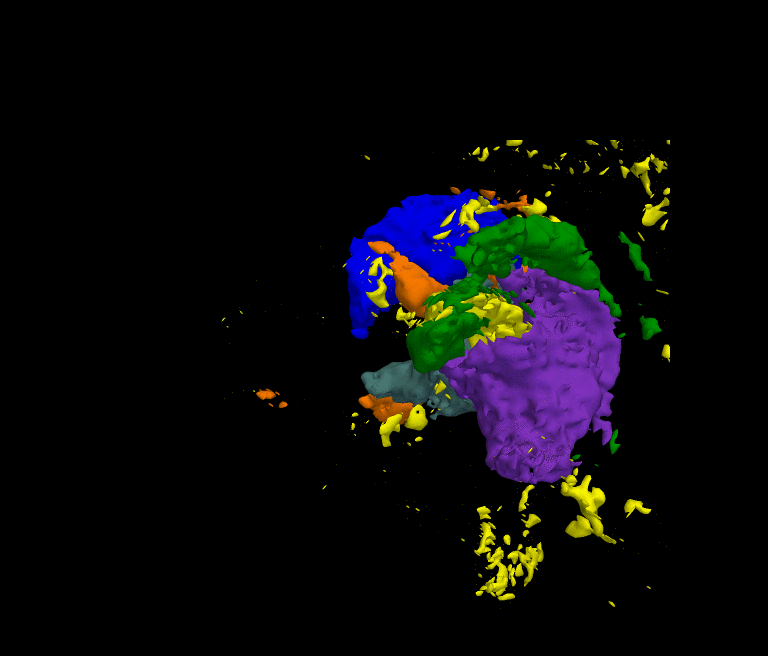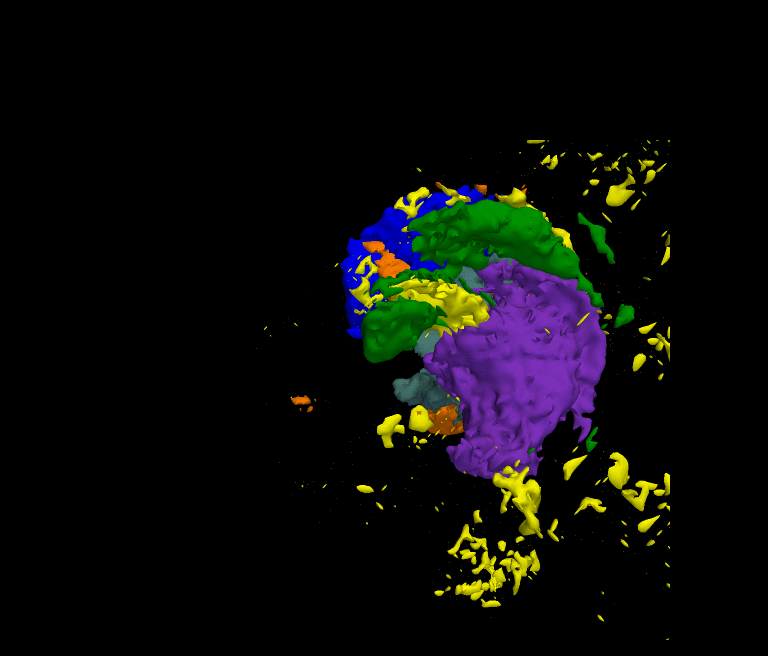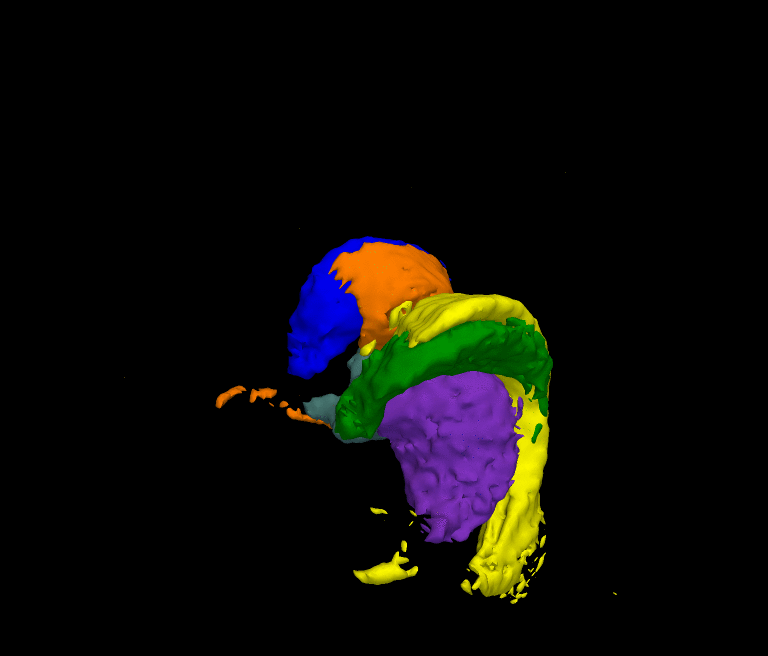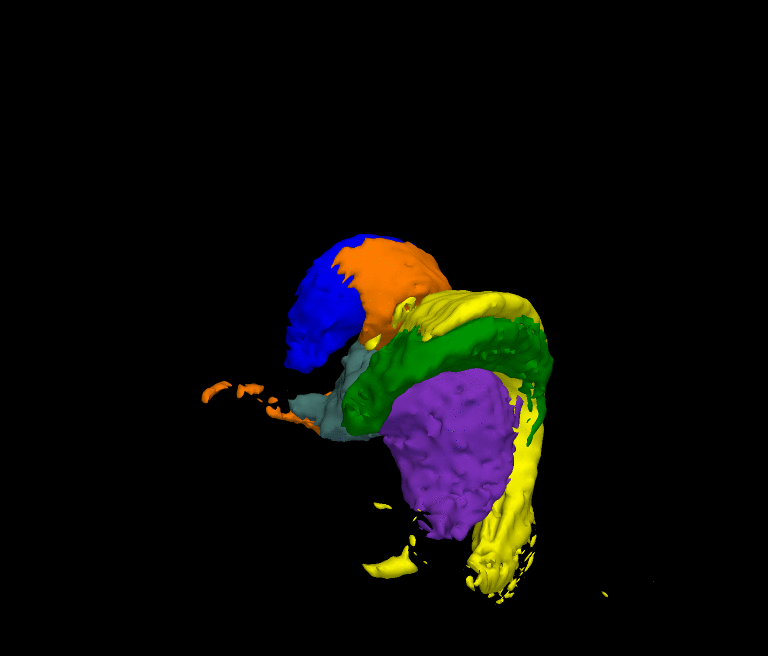
The movie shows the segmentation results of our approach after each iteration. The thalamus is represented by dark green and purple, the caudate by blue and light green, and the ventricles by orange and yellow. The noisy initial segmentation is caused by the misalignment of the atlas to the image space and the incorrect deformation of the shape model. As the method progresses the quality of the segmentation improves.
Experiement of 22 Cases
|
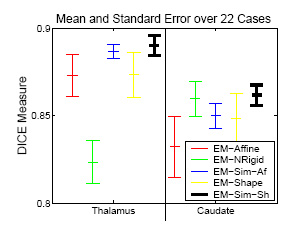
The experiment empirically demonstrates the utility of joining registration and shape based segmentation in an EM implementation by comparing the accuracy our new method with (EM-Sim-Sh) to three other EM implementations:
All methods segment 22 cases into the three brain tissue classes as well as the ventricles, thalamus, and caudate. We then measure the overlap between manual and automatic segmentations of the thalamus and caudate using DICE. Only our new approach (EM-Sim-Sh) performs well for both structures. |
Publications
[1] K.M. Pohl, J. Fisher, M. Shenton, R. W. McCarley, W.E.L. Grimson, R. Kikinis, and W.M. Wells. Logarithm odds maps for shape representation. In Proc. MICCAI 2006: Ninth International Conference Medical Image Computing and Computer-Assisted Intervention, Springer-Verlag, vol 4191 of Lecture Notes in Computer Science, pp 955–963, 2006
[2] K. M. Pohl, J. Fisher, W.E.L. Grimson, R. Kikinis, and W.M. Wells. A Bayesian Model for Joint Segmentation and Registration. NeuroImage, 31(1), pp. 228-239, 2006
[3] K.M. Pohl, J. Fisher, J.J. Levitt, M.E. Shenton, R. Kikinis, W.E.L. Grimson, and W.M. Wells: A Unifying Approach to Registration, Segmentation, and Intensity Correction, In Proc. MICCAI’2005: Eigth International Conference on Medical Image Computing and Computer Assisted Intervention, 2005.
[4] K.M. Pohl, J. Fisher, R. Kikinis, W.E.L. Grimson, and W.M. Wells: Shape Based Segmentation of Anatomical Structures in Magnetic Resonance Images, In Proc. ICCV’2005: Computer Vision for Biomedical Image Applications: Current Techniques and Future Trend, An International Conference on Computer Vision Workshop, 2005.
[5] K.M. Pohl: Prior Information for Brain Parcellation, PhD-Thesis, Department of Electrical Engineering and Computer Science, Massachusetts Institute of Technology, 2005.
[6] K.M. Pohl, John Fisher, W. Eric L. Grimson, William M. Wells: An Expectation Maximization Approach for Integrated Registration, Segmentation, and Intensity Correction CSAIL Publications - Artificial Intelligence Series Publications, AIM-2005-010, pp. 1-13, 2005.
[7] K.M. Pohl, S.K. Warfield, R. Kikinis, W.E. L. Grimson, W.M. Wells, Coupling Statistical Segmentation and PCA Shape Modeling, In Proc. MICCAI 2004: Seventh International Conference on Medical Image Computing and Computer Assisted Intervention, Rennes / St-Malo, France, Springer-Verlag, vol. 3216 of Lecture Notes in Computer Science, pp. 151-159, 2004
[8] Michael Leventon, Eric Grimson, Olivier Faugeras. "Statistical Shape Influence in Geodesic Active Contours" Comp. Vision and Patt. Recon. (CVPR), 2000.
[9] Wells, W., Grimson, W., Kikinis, R., Jolesz, F., 1996. Adaptive segmentation of MRI data. IEEE Transactions in Medical Imaging 15, 429–442.
Software
The algorithm is currently implemented in 3D Slicer Version 2.6. We are working on integrating the implementation in 3D Slicer (Version 3) with a resigned and improved user interface.