Difference between revisions of "NA-MIC/Projects/Collaboration/MGH RadOnc"
| Line 1: | Line 1: | ||
__NOTOC__ | __NOTOC__ | ||
| − | |||
===Adaptive RT for Head and Neck=== | ===Adaptive RT for Head and Neck=== | ||
| Line 22: | Line 21: | ||
===Adaptive RT for Thorax=== | ===Adaptive RT for Thorax=== | ||
| − | This example shows | + | This example shows anatomic change in the thorax. The patient has a collapsed left lower lobe in the pre-treatment scan (left), which has recovered in the mid-treatment scan (right). Notice there is some kind of fluid accumulation below the collapsed lung. |
{| | {| | ||
| Line 29: | Line 28: | ||
|} | |} | ||
| − | Here is another | + | Here is another example for thorax. The tumor has regressed during treatment. |
{| | {| | ||
| Line 35: | Line 34: | ||
|[[Image:RadOnc_HN5_deformable.png|thumb|380px|Deformable registration]] | |[[Image:RadOnc_HN5_deformable.png|thumb|380px|Deformable registration]] | ||
|} | |} | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
Thorax is a special case. Patient images are acquired using 4D-CT, and radiation dose can computed for 3D volumes at each breathing phase. The volumes are aligned using deformable registration, and radiation dose is accumulated in a reference phase (e.g. exhale). Ideally this procedure is repeated to perform 4D treatment plan optimization. | Thorax is a special case. Patient images are acquired using 4D-CT, and radiation dose can computed for 3D volumes at each breathing phase. The volumes are aligned using deformable registration, and radiation dose is accumulated in a reference phase (e.g. exhale). Ideally this procedure is repeated to perform 4D treatment plan optimization. | ||
| Line 60: | Line 53: | ||
State of the art is probably around 2-3 mm RMS error for point landmarks. | State of the art is probably around 2-3 mm RMS error for point landmarks. | ||
| − | + | ===General Discussion of Registration=== | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | Deformable registration is still not as reliable as it should be. Here are some of my complaints: | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | # Image acquisition has residual artifacts which cause unrealistic deformations (dental artifacts in H&N, motion artifacts in thorax). | |
| + | # Registration algorithms are not always robust, and require experimentation and tuning. | ||
| + | # Validation of registration results is not easy, since there are inadequate tools. | ||
| + | # In thorax, temporal regularization is generally not done, because of slow algorithms and large memory footprints. | ||
| + | # For cone-beam CT (and sometimes MR), intensity values are not globally stable. The most common suggestion is adaptive histogram equalization, but isn't there a better way? | ||
| + | # Interactive tools to repair (or reinitialize) registration are virtually non-existent. | ||
| − | + | ===Segmentation=== | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | == | ||
Rad Onc departments use interactive segmentation every day for both research and patient care. | Rad Onc departments use interactive segmentation every day for both research and patient care. | ||
| Line 99: | Line 76: | ||
# post-processing tools to nudge or smooth the boundary | # post-processing tools to nudge or smooth the boundary | ||
| − | There are many opportunities for using computer vision algorithms to improve interactive segmentation. For example, using prior models of shape and intensity to improve interpolation. | + | There are many opportunities for using computer vision algorithms to improve interactive segmentation. For example, using prior models of shape and intensity to improve interpolation. |
| + | |||
| + | In addition, the here are several commercial products for automatic segmentation, each with impressive demos. Personally I have never used any of them, but I hear that segmentation of simple structures are already useful (spinal cord, lens & optic nerve, liver). | ||
Revision as of 21:01, 29 January 2009
Home < NA-MIC < Projects < Collaboration < MGH RadOnc
Adaptive RT for Head and Neck
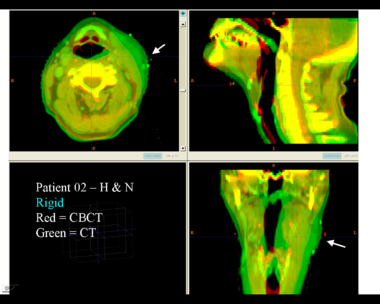
This first example shows the kind of anatomic changes that can occur in head and neck cancer. The pre-treatment scan is in green, and the mid-treatment scan is in red. The image on the left is the rigid registration, on the right is deformable registration. The deformable registration is
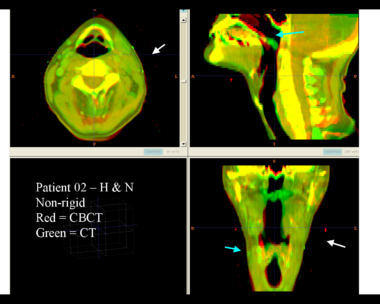
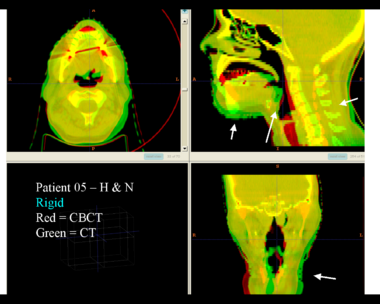
Here is another pertinent example for head and neck. In axial view, there appears to be some weight loss. Note the change in positioning of the mandible, and also the twisting of the cervical spine between scans. Also note the strong CT artifacts caused by dental fillings. In both examples, registration of the soft palette is worse using deformable registration than rigid registration, probably due to these artifacts.
| File:RadOnc HN5 deformable.png |
(Why the wiki doesn't display the thumbnail for the right image? Anyone?)
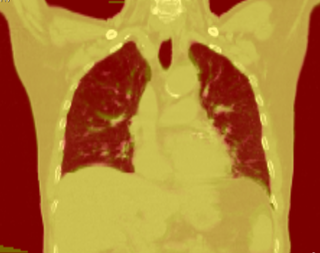
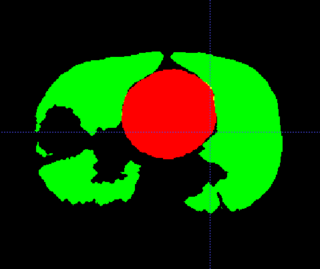
Adaptive RT for Thorax
This example shows anatomic change in the thorax. The patient has a collapsed left lower lobe in the pre-treatment scan (left), which has recovered in the mid-treatment scan (right). Notice there is some kind of fluid accumulation below the collapsed lung.
Here is another example for thorax. The tumor has regressed during treatment.
| File:RadOnc HN5 deformable.png |
Thorax is a special case. Patient images are acquired using 4D-CT, and radiation dose can computed for 3D volumes at each breathing phase. The volumes are aligned using deformable registration, and radiation dose is accumulated in a reference phase (e.g. exhale). Ideally this procedure is repeated to perform 4D treatment plan optimization.
The sliding of the lungs against the chest wall is difficult to model. We sometimes segment the images at the pleural boundary. This allows us to separate the moving set of organs from the non-moving set, which are registered separately. Ideally we would always do this, but segmentation is manual and therefore we usually skip this step.
If you ignore the pleural boundary, registration of 4D-CT is considered "easy".
- Single-session imaging, so patient is already co-registered
- Single-session imaging, so no anatomic change
- High contrast of vessels against lung parenchema
State of the art is probably around 2-3 mm RMS error for point landmarks.
General Discussion of Registration
Deformable registration is still not as reliable as it should be. Here are some of my complaints:
- Image acquisition has residual artifacts which cause unrealistic deformations (dental artifacts in H&N, motion artifacts in thorax).
- Registration algorithms are not always robust, and require experimentation and tuning.
- Validation of registration results is not easy, since there are inadequate tools.
- In thorax, temporal regularization is generally not done, because of slow algorithms and large memory footprints.
- For cone-beam CT (and sometimes MR), intensity values are not globally stable. The most common suggestion is adaptive histogram equalization, but isn't there a better way?
- Interactive tools to repair (or reinitialize) registration are virtually non-existent.
Segmentation
Rad Onc departments use interactive segmentation every day for both research and patient care. Prior to treatment planning, the target and critical structures are delineated in CT. The current state of the art is manual segmentation in axial view. A outline tool, used delineate the boundary, is generally prefered over a paintbrush tool that fills pixels. Commercial products generally support some subset of the following tools to assist the operator.
- contour interpolation between slices
- boundary editing
- mixed axial/coronal/sagittal drawing
- livewire or intelligent scissors
- drawing constraints (e.g. constraints on volume overlap/distance)
- post-processing tools to nudge or smooth the boundary
There are many opportunities for using computer vision algorithms to improve interactive segmentation. For example, using prior models of shape and intensity to improve interpolation.
In addition, the here are several commercial products for automatic segmentation, each with impressive demos. Personally I have never used any of them, but I hear that segmentation of simple structures are already useful (spinal cord, lens & optic nerve, liver).