Unused files
From NAMIC Wiki
The following files exist but are not embedded in any page. Please note that other web sites may link to a file with a direct URL, and so may still be listed here despite being in active use.
Showing below up to 500 results in range #501 to #1,000.
View (previous 500 | next 500) (20 | 50 | 100 | 250 | 500)
- Algorithms-Logo.png 75 × 75; 7 KB
- Engineering-Logo.png 75 × 75; 8 KB
- Dbp-logo.png 75 × 75; 8 KB
- Service-Logo.png 75 × 75; 7 KB
- Training-Logo.png 75 × 75; 8 KB
- Dissemination-Logo.png 75 × 74; 7 KB
- Management-Logo.png 75 × 74; 7 KB
- ToolbarMouseLasso.png 21 × 21; 487 bytes
- ToolbarMouseLassoLow.png 21 × 21; 499 bytes
- SlicerToolbarIcons2.png 782 × 1,152; 96 KB
- IconBlank.png 21 × 21; 248 bytes
- IconBlankLow.png 21 × 21; 249 bytes
- MenuButtonIconBlank.png 21 × 21; 286 bytes
- MenuButtonIconBlankDisabled.png 21 × 21; 300 bytes
- MayaInterface.png 987 × 686; 97 KB
- MouseModeMockup2.png 773 × 587; 20 KB
- MouseModeIcons.png 236 × 58; 6 KB
- Tetmesh namic.ppt ; 258 KB
- MouseModeMockup.png 720 × 936; 77 KB
- SB1.png 506 × 542; 123 KB
- SB3.png 505 × 529; 148 KB
- SB4.png 506 × 543; 188 KB
- SB5.png 505 × 543; 189 KB
- SB6.png 506 × 542; 190 KB
- SB7.png 506 × 543; 188 KB
- SB8.png 506 × 543; 191 KB
- SB9.png 507 × 556; 191 KB
- SB10.png 506 × 556; 191 KB
- SB2.png 506 × 542; 159 KB
- Test-Image-Logo.png 75 × 74; 7 KB
- KarssenbergBridge02-mj.pdf 0 × 0; 1.18 MB
- Margulies-mj.pdf 0 × 0; 913 KB
- Margulies-mj.png 456 × 274; 130 KB
- Carlson-mj.pdf 0 × 0; 164 KB
- Carlson-mj.png 430 × 600; 201 KB
- Royal-society-mj.pdf 0 × 0; 459 KB
- Akselrod-ballin-mj.pdf 0 × 0; 1.94 MB
- Zeevi-mj.jpg 512 × 420; 15 KB
- Deisboeck.jpg 612 × 274; 29 KB
- Mongkolwat.jpg 481 × 600; 42 KB
- Li-ClinicalAnatomy2006-mj.pdf 0 × 0; 204 KB
- Li-ClinicalAnatomy2006-fig-mj.png 650 × 286; 209 KB
- Group-Slicer2006-mj.pdf 0 × 0; 1.15 MB
- Group-Slicer2006-fig-mj.png 567 × 346; 252 KB
- SlicesFitToWindow.png 21 × 21; 3 KB
- Jason & Lara.txt ; 1,011 bytes
- DwiTstCase.tar.gz ; 23.22 MB
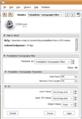
- CMake-Python1.png 679 × 452; 22 KB
- PythonMenu.png 390 × 296; 27 KB
- PythonConsole.png 789 × 500; 51 KB
- Slicer2.6-opt-win32-x86-2007-02-11.zip ; 36.92 MB
- Splot.zip ; 3.85 MB
- Skull-ct.zip ; 212 KB
- PythonBefore.png 918 × 393; 146 KB
- PythonAfter.png 919 × 397; 90 KB
- Pythonimshow.png 681 × 582; 118 KB
- Pythonhist.png 681 × 583; 29 KB
- Image info.jpg 981 × 725; 108 KB
- P41 retreat 07-01-29.ppt ; 9.65 MB
- Farke-Morphology2007-fig-mj.jpg 797 × 833; 167 KB
- Farke-Morphology2007-mj.pdf 0 × 0; 400 KB
- Farke-Paleontology2006-fig-mj.png 1,063 × 633; 157 KB
- Farke-Paleontology2006-fig-mj.jpg 797 × 833; 167 KB
- Farke-Paleontology2006-mj.pdf 0 × 0; 320 KB
- FiducialNavigationSliceWindow.png 308 × 365; 8 KB
- Splot2.zip ; 3.72 MB
- Murphy Workflow AHM 2007-Wed.ppt ; 3.35 MB
- Astromed1-mj.jpg 486 × 196; 16 KB
- NavigationWindow.png 1,318 × 976; 492 KB
- ZoomWindow.png 1,318 × 976; 438 KB
- Sensor-IntegratedVRforSurgicalApps-mj.pdf 0 × 0; 2.11 MB
- King-mj.png 523 × 408; 199 KB
- Kotsas-mj.png 359 × 189; 32 KB
- Royal-society-mj.png 444 × 468; 235 KB
- Neuroimage22,720.png 463 × 444; 181 KB
- Neuroimage22,720-mj.png 463 × 444; 181 KB
- Chen-mj.png 644 × 237; 91 KB
- Akselrod-ballin-mj.png 642 × 305; 202 KB
- Dierks-99-lateral.gif 426 × 186; 45 KB
- Lennox-2000-all.gif 432 × 181; 46 KB
- Table1.PNG 623 × 131; 9 KB
- Table2.PNG 603 × 147; 10 KB
- Curv-axial.png 75 × 75; 3 KB
- Curv-coronal.png 75 × 75; 3 KB
- Curv-saggital.png 75 × 75; 3 KB
- Cbg-fa-axial.png 75 × 75; 3 KB
- Cbg-fa-coronal.png 75 × 75; 2 KB
- Cbg-fa-saggital.png 75 × 75; 3 KB
- Shergill-2004-all.jpg 429 × 187; 58 KB
- Kubicki 2005NYAS.pdf 0 × 0; 3.69 MB
- Kubicki 2005NYAS-mj.png 680 × 391; 373 KB
- StructureImprovementOverFreesurferLateral.jpg 1,131 × 721; 184 KB
- StructureImprovementOverFreesurferMedial.jpg 1,131 × 659; 195 KB
- Butaric-slicer-mj.pdf 0 × 0; 428 KB
- Butaric-mj.png 418 × 299; 79 KB
- DSC00838.JPG 3,072 × 2,304; 2.71 MB
- Portal-main.png 830 × 758; 256 KB
- Portal-logged-in.png 967 × 788; 232 KB
- Projects-tab.png 967 × 788; 326 KB
- Membership-request.png 936 × 274; 117 KB
- Membership-request-accepted.png 957 × 329; 174 KB
- Namic-project.png 1,009 × 301; 83 KB
- Search-icon.gif 48 × 48; 3 KB
- Search-icon.png 48 × 48; 3 KB
- Namic-project-space.png 967 × 788; 307 KB
- Namic-file-space.png 953 × 481; 106 KB
- Slicer 3-beta-2007-04-16.ppt ; 266 KB
- SerdarAffineOrig.jpg 256 × 256; 6 KB
- SerdarAffineRegister.jpg 256 × 256; 6 KB
- SerdarBSplineOrig.jpg 256 × 256; 5 KB
- SerdarBSplineRegister.jpg 256 × 256; 6 KB
- MeanOriginal.png 256 × 256; 12 KB
- MeanRegisteredAffine.png 256 × 256; 9 KB
- MeanRegisteredBspline.png 256 × 256; 9 KB
- CingulumPrototype.png 334 × 153; 798 bytes
- Ipmi07-pohl.pdf 0 × 0; 854 KB
- Rocket.jpg 127 × 127; 7 KB
- Slicer-launch.png 1,062 × 526; 122 KB
- Slicer-launched.png 510 × 314; 27 KB
- 180px-Lcomb-grayscale.png 180 × 135; 30 KB
- 180px-Meanviews.png 180 × 143; 79 KB
- Slicer3interface.png 417 × 606; 19 KB
- Pieper-MGH-medical-physics-2007-04-24.ppt ; 31.13 MB
- Cascio-Am-J-Psych-163-12-Dec06.pdf 0 × 0; 198 KB
- 5bundles.jpg 631 × 565; 63 KB
- FAs.jpg 594 × 345; 46 KB
- Slide1.jpg 793 × 595; 75 KB
- SlicerAstrocyte.jpg 793 × 595; 75 KB
- NCMIR example dataset.zip ; 107.82 MB
- Pohl-mj.jpg 311 × 305; 16 KB
- Svn-macosx-ssl-static.zip ; 2.42 MB
- Pohl CT3DSegmentation.jpg 455 × 444; 23 KB
- 3DVisualization NCIA SPujol.pdf 0 × 0; 1.51 MB
- 2007 ISMRM GE.zip ; 89.66 MB
- Spl pnl brain atlas 2007.zip ; 32.06 MB
- SPL-BrainAtlas-label list.doc ; 151 KB
- Spl pnl brain atlas 2007a.zip ; 32.06 MB
- Recv data.xml.txt ; 446 bytes
- Send data.xml.txt ; 463 bytes
- NTTutorial May2007.tar.gz ; 2 KB
- NAC Poster mBIRN Collaboration.ppt ; 939 KB
- VmtkSlicerModule.png 397 × 270; 58 KB
- NTTutorial May2007 2.zip ; 36 KB
- Na-mic core1 200705 meeting lupus progress report.ppt.pdf 0 × 0; 1.41 MB
- Atlas-2007-05-16.png 2,469 × 1,578; 1.07 MB
- CineDisplayDesigns.png 1,728 × 390; 32 KB
- Cortex-Thick.ppt ; 5.56 MB
- Slicer2.6-LE-opt-win32-x86-2006-12-08.zip ; 33.37 MB
- Sample DTI.zip ; 14.81 MB
- Slicer2.6-opt-win32-x86-2006-05-19-old.zip ; 31.77 MB
- Kwmeshvisu-slicer-logo.png 1,280 × 744; 65 KB
- EMSegmentTutorial Slicer3.zip ; 41.23 MB
- EMSegmentTutorial.tgz ; 41.07 MB
- Dti namic yendiki.ppt ; 1.83 MB
- Intro.DTI.Workshop.OHBM.2007.ppt ; 21.55 MB
- Intro-DTI-Workshop-OHBM2007.ppt ; 21.55 MB
- Cobb-BiomedicalComputationReview2007.pdf 0 × 0; 616 KB
- Cobb-BiomedicalComputationReview2007-fig.png 1,434 × 504; 245 KB
- Statementofintent.doc ; 28 KB
- Statementofintent.pdf 0 × 0; 49 KB
- BWHstatementofintent.pdf 0 × 0; 89 KB
- Slicer3 DTI tractographyDevel 1.png 700 × 466; 193 KB
- 3DSlicerLogo-DesktopIcon-16x16x16.gif 16 × 16; 223 bytes
- 3DSlicerLogo-DesktopIcon-16x16x16.png 16 × 16; 419 bytes
- 3DSlicerLogo-DesktopIcon-16x16x256.gif 16 × 16; 1 KB
- 3DSlicerLogo-DesktopIcon-16x16x256.png 16 × 16; 948 bytes
- 3DSlicerLogo-DesktopIcon-32x32x16.gif 32 × 32; 504 bytes
- 3DSlicerLogo-DesktopIcon-32x32x16.png 32 × 32; 838 bytes
- 3DSlicerLogo-DesktopIcon-32x32x256.gif 32 × 32; 2 KB
- 3DSlicerLogo-DesktopIcon-32x32x256.png 32 × 32; 2 KB
- 3DSlicerLogo-DesktopIcon-48x48x16.gif 48 × 48; 884 bytes
- 3DSlicerLogo-DesktopIcon-48x48x16.png 48 × 48; 1 KB
- 3DSlicerLogo-DesktopIcon-48x48x256.gif 48 × 48; 3 KB
- 3DSlicerLogo-DesktopIcon-48x48x256.png 48 × 48; 3 KB
- 3DSlicerLogo-DesktopIcon-64x64x16.gif 64 × 64; 1 KB
- 3DSlicerLogo-DesktopIcon-64x64x16.png 64 × 64; 2 KB
- 3DSlicerLogo-DesktopIcon-64x64x256.gif 64 × 64; 4 KB
- 3DSlicerLogo-DesktopIcon-64x64x256.png 64 × 64; 4 KB
- 3DSlicerLogo-DesktopIcon-128x128x16.gif 128 × 128; 4 KB
- 3DSlicerLogo-DesktopIcon-128x128x16.png 128 × 128; 4 KB
- 3DSlicerLogo-DesktopIcon-128x128x256.gif 128 × 128; 8 KB
- 3DSlicerLogo-DesktopIcon-128x128x256.png 128 × 128; 7 KB
- Slicer3MyModule.png 779 × 954; 158 KB
- DtiDevel2.jpg 800 × 527; 218 KB
- Slicer3Palette.png 288 × 113; 5 KB
- DiffusionTODOList.jpg 640 × 480; 88 KB
- ArcuateROI.png 512 × 512; 127 KB
- Fornix1ROIs.png 512 × 512; 98 KB
- Fornix2ROIs.png 512 × 512; 94 KB
- KWDirectoryBrowserLinuxTooShort.jpg 510 × 404; 35 KB
- KWMeshViewer030607.png 840 × 658; 82 KB
- Macaque.png 135 × 234; 59 KB
- Brainseg1.jpg 753 × 563; 52 KB
- BeforePicture.JPG 821 × 331; 46 KB
- BeforePicture.jpg 821 × 331; 46 KB
- AddPositionVertex.png 21 × 21; 356 bytes
- ChooseBrushSize.png 21 × 21; 543 bytes
- ChooseColor.png 21 × 21; 264 bytes
- Close.png 21 × 21; 333 bytes
- ConvertROI2Label.png 21 × 21; 340 bytes
- CopyROI.png 21 × 21; 345 bytes
- DefineAndEditROIs.png 21 × 21; 392 bytes
- Delete.png 21 × 21; 349 bytes
- Deselect.png 21 × 21; 317 bytes
- DilateROI.png 21 × 21; 577 bytes
- DrawExplicitPolylineAnnularROI.png 21 × 21; 384 bytes
- DrawExplicitPolylineROI.png 21 × 21; 411 bytes
- EraseLabel.png 21 × 21; 423 bytes
- ErodeROI.png 21 × 21; 593 bytes
- GoToEditor.png 21 × 21; 426 bytes
- ImplicitCubeROI.png 21 × 21; 347 bytes
- ImplicitEllipseROI.png 21 × 21; 334 bytes
- ImplicitRectangleROI.png 21 × 21; 300 bytes
- ImplicitSphereROI.png 21 × 21; 487 bytes
- InterpolateROIs.png 21 × 21; 382 bytes
- MakeModel.png 21 × 21; 454 bytes
- MoveVertex.png 21 × 21; 359 bytes
- PaintLabel.png 21 × 21; 496 bytes
- PasteROI.png 21 × 21; 356 bytes
- PinOpen.png 21 × 21; 611 bytes
- SelectROI.png 21 × 21; 372 bytes
- SelectVertex.png 21 × 21; 380 bytes
- SetROIOpacity.png 21 × 21; 390 bytes
- SlurpColor.png 21 × 21; 393 bytes
- SnapToGridOff.png 21 × 21; 372 bytes
- SnaptoGridOn.png 21 × 21; 331 bytes
- TrimROI.png 21 × 21; 384 bytes
- UnionROI.png 21 × 21; 367 bytes
- DrawFreePolylineROI.png 21 × 21; 447 bytes
- DrawFreePolylineAnnularROI.png 21 × 21; 346 bytes
- RedrawFreePolylineROI.png 21 × 21; 368 bytes
- Continuous-build.png 613 × 190; 10 KB
- SelectAllGuides.png 21 × 21; 327 bytes
- DrawFreehandGuide.png 21 × 21; 366 bytes
- DrawExplicitPolylineGuide.png 21 × 21; 301 bytes
- ConvertGuide2ROI.png 21 × 21; 344 bytes
- After.PNG 1,306 × 2,131; 170 KB
- After.png 1,306 × 2,131; 170 KB
- LightBoxProjectWeek2007.png 1,024 × 744; 2.18 MB
- LightBoxProjectWeek2007 blackBackground.png 1,024 × 768; 2.25 MB
- DiffusionImagingMRML.png 2,134 × 1,122; 30 KB
- SelectFiducial.png 21 × 21; 318 bytes
- PrevFiducial.png 21 × 21; 357 bytes
- NextFiducial.png 21 × 21; 355 bytes
- Fiducial.png 21 × 21; 272 bytes
- SlicerVisComm.png 600 × 234; 37 KB
- RadioIcons.png 259 × 969; 24 KB
- SmallEmbodiment.png 600 × 464; 83 KB
- SmallSliceGUI.png 400 × 262; 65 KB
- RadioCheckButtons.png 535 × 725; 20 KB
- Slicer3-xnat.png 749 × 856; 380 KB
- MouseModeToolbar.png 432 × 242; 11 KB
- EditorModuleDesign.png 842 × 767; 47 KB
- Li-ClinAnat2006.pdf 0 × 0; 204 KB
- Li-ClinAnat2006-mj.png 472 × 267; 144 KB
- Hunter-Rheumatology2003.pdf 0 × 0; 106 KB
- Hunter-OsteoArthritisCartilage2003.pdf 0 × 0; 90 KB
- Slicer3-coordinates.png 1,123 × 845; 52 KB
- DeselectedRadioCheckButtonIconBlank.png 21 × 21; 248 bytes
- DisabledRadioCheckButtonIconBlank.png 21 × 21; 256 bytes
- BottomSelectedCheckIndicatorAndIconBlank.png 21 × 36; 414 bytes
- BottomDeselectedIndicatorAndIconBlank.png 21 × 36; 354 bytes
- BottomDisabledIndicatorAndIconBlank.png 21 × 36; 373 bytes
- BottomSelectedRadioIndicatorAndIconBlank.png 21 × 36; 390 bytes
- TopSelectedCheckIndicatorAndIconBlank.png 21 × 36; 413 bytes
- LeftSelectedCheckIndicatorAndIconBlank.png 36 × 21; 375 bytes
- RightSelectedCheckIndicatorAndIconBlank.png 36 × 21; 360 bytes
- TopDeselectedIndicatorAndIconBlank.png 21 × 36; 357 bytes
- TopDisabledIndicatorAndIconBlank.png 21 × 36; 372 bytes
- LeftDeselectedIndicatorAndIconBlank.png 36 × 21; 306 bytes
- LeftDisabledIndicatorAndIconBlank.png 36 × 21; 336 bytes
- RightDeselectedIndicatorAndIconBlank.png 36 × 21; 290 bytes
- RightDisabledIndicatorAndIconBlank.png 36 × 21; 336 bytes
- TopSelectedRadioIndicatorAndIconBlank.png 21 × 36; 392 bytes
- LeftSelectedRadioIndicatorAndIconBlank.png 36 × 21; 336 bytes
- RightSelectedRadioIndicatorAndIconBlank.png 36 × 21; 328 bytes
- CheckIndicatorOn.png 11 × 11; 264 bytes
- RadioIndicatorOn.png 11 × 11; 237 bytes
- IndicatorOff.png 11 × 11; 212 bytes
- IndicatorDisabled.png 11 × 11; 212 bytes
- LikelihoodsFiberResultOverBaseline018.png 488 × 276; 15 KB
- SlicerIcons.png 728 × 484; 47 KB
- Forn roi1.png 512 × 512; 163 KB
- Forn roi2.png 512 × 512; 159 KB
- MonkeyDTIStudy Neuroimage2007.pdf 0 × 0; 912 KB
- BrainTissueClassifiers BouixNeuroimage2007.pdf 0 × 0; 3.55 MB
- ROI1 ax0001.jpg 512 × 512; 51 KB
- ROI1 sag.jpg 512 × 512; 41 KB
- ROI2 cor.jpg 512 × 512; 45 KB
- ROI2 sagL.jpg 512 × 512; 51 KB
- ROI2 sagR.jpg 512 × 512; 52 KB
- Editbox.png 532 × 439; 110 KB
- Software-plan.ppt ; 55 KB
- EditorModuleDesignGen1.png 842 × 720; 46 KB
- Gen1EditorSketch.png 576 × 492; 22 KB
- Livers.png 949 × 986; 116 KB
- MRI Robot System Diagram.png 809 × 606; 26 KB
- Significance testing.jpg 856 × 862; 104 KB
- Screenshot-VNC-X11.png 1,290 × 1,052; 72 KB
- VNCscreenshot.png 1,290 × 1,052; 72 KB
- 2007 ISMRM IGT Plenary.zip ; 132.65 MB
- VNC2.png 1,290 × 1,052; 82 KB
- Stanford.ppt ; 55.89 MB
- Address.pdf 0 × 0; 19 KB
- Slicer3SampleVisualization.tar.gz ; 15.31 MB
- VolumeRenderingWithShading.jpg 758 × 568; 47 KB
- VolumeRendering.jpg 840 × 472; 68 KB
- VolumeRenderingBoneDetection.png 1,322 × 976; 384 KB
- Martinos logo big.jpg 624 × 668; 21 KB
- LONI Logo Head.gif 190 × 210; 17 KB
- Firdaus image 004.png 800 × 600; 30 KB
- Firdaus affine rms.png 800 × 600; 30 KB
- Firdaus affine mi.png 800 × 600; 29 KB
- Test1.doc ; 24 KB
- Original.jpg 1,211 × 1,037; 79 KB
- Resampled.jpg 1,211 × 1,037; 72 KB
- Affine-rms-registered.jpg 1,211 × 1,037; 79 KB
- Bspline-demo.mov ; 52.99 MB
- UseCaseVolumeRendering.jpg 1,236 × 676; 94 KB
- SlicerReformat.png 1,400 × 1,016; 259 KB
- GUI.jpg 751 × 1,115; 141 KB
- ClassDiagramVolumeRendering.jpg 1,166 × 656; 65 KB
- Picturetest.ppt ; 10 KB
- RicianTensorCorrectionImage.png 577 × 946; 1.12 MB
- Slicer-eng-u41.ppt ; 1.57 MB
- Slicer igt img1.jpg 512 × 384; 35 KB
- Slicer igt img2.jpg 512 × 384; 25 KB
- Slicer igt img3.jpg 512 × 384; 24 KB
- Slicer igt img4.jpg 512 × 384; 33 KB
- Slicer igt img5.jpg 512 × 384; 36 KB
- Slicer igt img6.jpg 512 × 384; 35 KB
- QA Presentation.zip ; 25.13 MB
- 936 fx.png 512 × 512; 138 KB
- Spl pnl brain atlas 2007 slicer3.zip ; 31.71 MB
- OUTPUT ic0.PNG 400 × 400; 9 KB
- OUTPUT ic1.PNG 400 × 400; 8 KB
- OUTPUT ic2.PNG 400 × 400; 9 KB
- OUTPUT ic3.PNG 400 × 400; 9 KB
- OUTPUT ic4.PNG 400 × 400; 8 KB
- OUTPUT ic5.PNG 400 × 400; 10 KB
- OUTPUT ic6.PNG 400 × 400; 9 KB
- OUTPUT ic7.PNG 400 × 400; 8 KB
- OUTPUT ic8.PNG 400 × 400; 9 KB
- OUTPUT ic9.PNG 400 × 400; 9 KB
- OUTPUT ic10.PNG 400 × 400; 8 KB
- OUTPUT ic11.PNG 400 × 400; 9 KB
- OUTPUT cing0.PNG 400 × 400; 9 KB
- OUTPUT cing1.PNG 400 × 400; 8 KB
- OUTPUT cing2.PNG 400 × 400; 9 KB
- OUTPUT cing3.PNG 400 × 400; 9 KB
- OUTPUT cing4.PNG 400 × 400; 8 KB
- OUTPUT cing5.PNG 400 × 400; 9 KB
- OUTPUT cing6.PNG 400 × 400; 8 KB
- OUTPUT cing7.PNG 400 × 400; 8 KB
- OUTPUT cing8.PNG 400 × 400; 9 KB
- OUTPUT cing9.PNG 400 × 400; 9 KB
- OUTPUT cing10.PNG 400 × 400; 8 KB
- OUTPUT cing11.PNG 400 × 400; 9 KB
- OUTPUT fx1.PNG 400 × 400; 8 KB
- OUTPUT fx0.PNG 400 × 400; 8 KB
- OUTPUT fx2.PNG 400 × 400; 9 KB
- OUTPUT fx3.PNG 400 × 400; 9 KB
- OUTPUT fx4.PNG 400 × 400; 9 KB
- OUTPUT fx5.PNG 400 × 400; 10 KB
- OUTPUT fx6.PNG 400 × 400; 8 KB
- OUTPUT fx8.PNG 400 × 400; 9 KB
- OUTPUT fx7.PNG 400 × 400; 8 KB
- OUTPUT fx9.PNG 400 × 400; 8 KB
- OUTPUT fx10.PNG 400 × 400; 9 KB
- OUTPUT fx11.PNG 400 × 400; 10 KB
- OUTPUT unc0.PNG 400 × 400; 9 KB
- OUTPUT unc1.PNG 400 × 400; 8 KB
- OUTPUT unc2.PNG 400 × 400; 9 KB
- OUTPUT unc3.PNG 400 × 400; 9 KB
- OUTPUT unc4.PNG 400 × 400; 8 KB
- OUTPUT unc5.PNG 400 × 400; 10 KB
- OUTPUT unc6.PNG 400 × 400; 9 KB
- OUTPUT unc7.PNG 400 × 400; 8 KB
- OUTPUT unc8.PNG 400 × 400; 8 KB
- OUTPUT unc9.PNG 400 × 400; 9 KB
- OUTPUT unc10.PNG 400 × 400; 8 KB
- OUTPUT unc11.PNG 400 × 400; 10 KB
- OUTPUT IC stats.xls ; 91 KB
- TractMeasures ic.xls ; 33 KB
- TractMeasures cingulum.xls ; 35 KB
- Slicer igt img7.jpg 512 × 384; 21 KB
- Slicer igt img8.jpg 512 × 384; 30 KB
- Slicer igt img9.jpg 512 × 384; 35 KB
- Slicer igt img10.jpg 512 × 384; 28 KB
- Slicer igt img11.jpg 512 × 384; 30 KB
- SantaFeTractography Gouttard.ppt ; 5.96 MB
- Goodlett-santa-fe-presentation.pdf 0 × 0; 4.71 MB
- QueryAtlasExampleData.zip ; 8.39 MB
- Slicer3QDEC AHM2007.ppt ; 1.99 MB
- QAloadQdec.png 1,323 × 909; 175 KB
- OntologyMapping.png 1,323 × 909; 416 KB
- QAqdecSearch.png 1,323 × 909; 171 KB
- QAsearch.png 1,323 × 909; 484 KB
- QAOntologyMapping.png 1,323 × 909; 416 KB
- FFSGUI.png 397 × 254; 9 KB
- QdecGUI.png 403 × 157; 6 KB
- DispGUI.png 404 × 104; 4 KB
- MappingGUI.png 405 × 366; 12 KB
- SearchGUI.png 399 × 219; 5 KB
- ResultsGUI.png 399 × 387; 13 KB
- ContextMenu.png 469 × 323; 108 KB
- Lexviewer.png 1,095 × 734; 59 KB
- QueryAtlas-AHM2007.zip ; 56.59 MB
- QueryAtlas-AHM2007.ppt ; 6.67 MB
- QAFIPSFreeSurfer.zip ; 56.59 MB
- QAQdec.zip ; 14.92 MB
- Xnat overview.gif 415 × 302; 17 KB
- ANN-Sagittal.gif 428 × 433; 32 KB
- DART-Logo.png 108 × 108; 6 KB
- Original image 004.png 768 × 545; 11 KB
- LiverRFAInitial.png 449 × 375; 72 KB
- IaFeMesh-small-102507.jpg 400 × 246; 9 KB
- Slicer-meshingworkflow-small-102507.jpg 400 × 279; 15 KB
- MappedMeshMiddleIndex.png 390 × 537; 63 KB
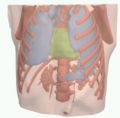
- AbdomenBones.jpg 1,600 × 1,200; 164 KB
- AbdomenSoftTisssue.jpg 1,600 × 1,200; 266 KB
- Python.zip ; 16 KB
- ICluster GM.gif 961 × 721; 94 KB
- ICluster GEM1.gif 961 × 721; 55 KB
- ICluster GEM2.gif 961 × 721; 47 KB
- ICluster GEM3.gif 961 × 721; 61 KB
- RobustnessOfWaveletAnalysis.png 1,327 × 349; 166 KB
- Mit fmri clustering parcellation3 shb1 4.png 493 × 609; 307 KB
- Gilmore.jpg 968 × 855; 329 KB
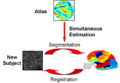
- Atlas demo.zip ; 76.69 MB
- Abdomen-v2.zip ; 24.1 MB
- MR intervention for dummies tempany 2007.ppt ; 48.99 MB
- NAMIC-SLC.tif 1,076 × 822; 2.54 MB
- SIGGRAPH-GPU.pdf 0 × 0; 6.39 MB
- LEGOIGTAndMedicalRoboticsTutorial 2 BasicTutorial.pdf 0 × 0; 2.56 MB
- LEGOIGTAndMedicalRoboticsTutorial 3 AdvancedTutorial.pdf 0 × 0; 2.2 MB
- Overcomplete vs biorthogonal wavelets.tif 1,362 × 582; 219 KB
- Slicer Architecture Discussion.png 589 × 180; 185 KB
- DwiGradientEditor.jpg 348 × 456; 35 KB
- DWI Gradient Editor.jpg 338 × 448; 35 KB
- EMSVisualizeTutorialInputData.png 1,218 × 695; 205 KB
- EMSVisualizeTutorialResultsData.png 1,218 × 695; 387 KB
- DTIUnregisted.jpg 536 × 447; 53 KB
- DTIRegisted.jpg 536 × 447; 55 KB
- DTIRgisted.jpg 536 × 447; 55 KB
- DTIregistration.jpg 521 × 340; 162 KB
- SlicerSampleVisualization.tar.gz ; 15.11 MB
- Volumerenderer.h.txt ; 146 KB
- DWI Gradient Editor v2.jpg 386 × 439; 129 KB
- RigidReg.jpg 234 × 103; 7 KB
- NonrigidReg.jpg 235 × 105; 8 KB
- DWI Gradient Editor v3.jpg 388 × 493; 147 KB
- Canine-heart1.png 1,248 × 1,109; 266 KB
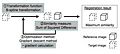
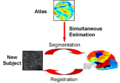
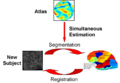
- JointRegSeg2.png 800 × 556; 135 KB
- JointRegSeg3.png 800 × 556; 135 KB
- Yeo-JointSeg-MICCAI2007.png 800 × 556; 137 KB
- CIMITlogo.gif 188 × 100; 7 KB
- Clustering small.png 200 × 171; 41 KB
- Heart256.raw ; 16 MB
- EMABC.jpg 200 × 125; 5 KB
- CellularReconstruction.png 538 × 1,011; 608 KB
- DuctRecon.png 517 × 579; 456 KB
- VtkCudaVolumeRenderingPipeline.png 805 × 622; 31 KB
- Slicer3 ProstateModule System Architecture.png 674 × 410; 29 KB
- Brain-seg-utah.tif 1,412 × 1,676; 109 KB
- HelloWorld.tar.gz ; 6.37 MB
- MattesGetValue.pdf 0 × 0; 36 KB
- MattesBSplineGetValue.pdf 0 × 0; 27 KB
- MeanSquaresGetValue.pdf 0 × 0; 30 KB
- MattesGetValueAndDerivative.pdf 0 × 0; 32 KB
- MattesBSplineGetValueAndDerivative.pdf 0 × 0; 29 KB
- MeanSquaresGetValueAndDerivative.pdf 0 × 0; 31 KB
- PythonModules.zip ; 13 KB
- SgmResult ghe100 withOriginal 113.png 746 × 1,006; 87 KB
- EMSegTutorial-AHM2008.zip ; 52.36 MB
- EMSegTutorial-AHM2008.ppt ; 2.76 MB
- Fiducial-seeding.jpg 1,439 × 874; 699 KB
- Roi tract.jpg 1,439 × 871; 735 KB
- Dti-color.jpg 1,440 × 869; 676 KB
- ENT.png 1,024 × 768; 71 KB
- FirstCudaVolumeRenderInSlicer3.png 1,201 × 460; 241 KB
- 2008 NA-MIC AHM Dissemination.ppt ; 1.37 MB
- NXT USB.zip ; 8 KB
- Ziyan-2007.jpg 458 × 522; 65 KB
- First4DVolumeRenderInSlicer3.png 1,173 × 786; 284 KB
- CMake-logo-med-res-c.png 120 × 118; 8 KB
- CineBasicController.png 356 × 75; 3 KB