Difference between revisions of "2011 Winter Project Week:Breakout Slicer with Ron"
From NAMIC Wiki
(→Data) |
|||
| Line 21: | Line 21: | ||
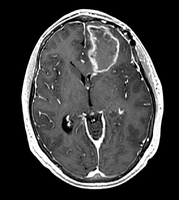
image:Case40-SPGRwGD.png|SPGR after injection of Gd | image:Case40-SPGRwGD.png|SPGR after injection of Gd | ||
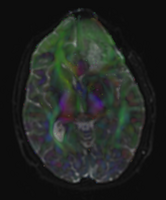
image:Case40-DTI.png|DTI color by orientation overlayed over T2 baseline | image:Case40-DTI.png|DTI color by orientation overlayed over T2 baseline | ||
| + | image:Case40Annotation.png|Annotation of some of the structures | ||
</gallery> | </gallery> | ||
Revision as of 15:39, 6 January 2011
Home < 2011 Winter Project Week:Breakout Slicer with RonBack to Project Week Agenda
Moderator: Ron Kikinis
Introduction
- This is an advanced topic presentation on Slicer abilities. The intention is to familiarize the NA-MIC community with Slicer abilities.
- You have mastered the introductions and you are hungry for more.... :)
- This means that we expect you to at least have worked through the Core Tutorials before this class.
- Ron and the creators of modules will show you advanced capabilities in Slicer 3.6
Logistics
- What: Advanced capabilities in Slicer
- Who: All the new DBP engineers, all the participants new to NA-MIC
- When: Monday afternoon: 3pm through 5pm
- Where: Main Event Room
Data
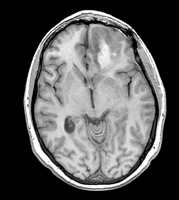
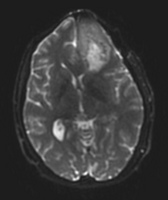
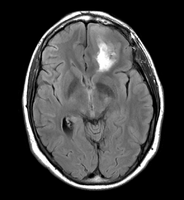
- Image data used
Case 40 from the fMRI neurosurgery data set on central.xnat.org
Preliminary Program
- 10 minutes or less per segment
- Comply with Rons Rules for Tools
- Registration (Dominik Meier, Hans Johnson)
- Register everything to the SPGR data: complete. Data, transforms and instructions available here: Reg.Lib Case 33
- additional info:
- basic concept and outline of DTI registration (cookbook) on FAQ, incl. List of special caveats pertaining to DTI registration and resampling
- overview of all DTI cases currently in the registration case library
- similar cases that required different approaches Case 29: DWI -> T2 -> T1 (direct to T1 fails), Case 03: low resol. & anisotropy require resampling, masking & manual initial alignment, Case 27: anisotropy required prior DWI resampling prior to conversion
- Resample as needed (DTI, T2 Baseline):
- transforms available here: Reg.Lib Case 33
- Compareview (Jim Miller)
- Scenesnapshots (Wendy Plesniak, Alex Yarmarkovich)
- Simple measurements and fiducials (Nicole Aucoin) Sample scene and data with snapshots for fiducials, seeding, measurements
- Fiducials tutorial and Data
- Adding and Deleting Fiducials
- Editing Fiducials
- Display properties
- Linking (control key+mouse move) and jumping slices (right click in table)
- Passing Fiducials to Command Line modules
- Fiducial Seeding
- Measurements
- Ruler (put down two fiducials, Control-m to make a ruler)
- Angle
- Constraining to slices or models
- Fiducials tutorial and Data
- Interactive editor (Steve Pieper)
- Fast segmenters: Fast Marching, RSS, GrowCutSegment (Andrey Fedorov, Yi Gao, Harini Veeraraghavan)
- Fast Marching demo scene
- inner part of the tumor segmented from SPGR
- outer part of the tumor segmented from post-Gad
- WM and GM segmented from N4-processed SPGR (WM segmentation of the original SPGR volume is included, note under-segmented WM in the skull base)
- all segmentations were done using FastMarching, fiducials are included for each of the segmentations
- no fine-tuning of the fiducial locations was done -- this is an example result one can get almost right away
- rule of thumb in placing fiducials: try to cover uniformly the volume you are trying to segment; this is particularly important for large structures like WM/GM
- RSS demo scene
- Segment the inner part of the tumor from SPGR using RSS
- Fast Marching demo scene
- DTI processing (Alex Yarmarkovich, Demian Wasserman)
- Volume Cropping (Andrey Fedorov)
- Keyboard and mouse shortcuts (Steve Pieper)
- Volume Rendering (Yanling Liu/Alex Yarmarkovich/Curtis Lisle)
- EM segmentation (Kilian Pohl)
Attendees
- Ron Kikinis
- Steve Pieper
- UCLA DBP Engineer: Andrei Irimia
- Iowa DBP Engineer: Mark Scully
- MGH DBP Engineer: Nadya Shusharina
- Utah DBP Engineer: Josh Cates
- Utah DBP Student: Josh Blauer
- Utah DBP PI: Rob MacLeod
- Utah DBP Student: Chris Gloschat