Difference between revisions of "Projects:RegistrationLibrary:RegLib C33"
From NAMIC Wiki
| Line 123: | Line 123: | ||
===Download === | ===Download === | ||
*Data | *Data | ||
| − | **[[Media:RegLib_C33_Data.zip |'''RegLib_C33_Data''' DWI, DTI, T1, FLAIR, Presets & solution transforms <small> (NRRD files, zip file | + | **[[Media:RegLib_C33_Data.zip |'''RegLib_C33_Data''' DWI, DTI, T1, FLAIR, Presets & solution transforms <small> (NRRD files, zip file 250 MB) </small>]]''' |
| + | **[[Media:RegLib_C33_Data_noDWI.zip |'''RegLib_C33_Data Subset''' as above, but w/o DWI and resampled DTI <small> (NRRD files, zip file 120 MB) </small>]]''' | ||
*Presets | *Presets | ||
**[[Media:RegLib_C33_Preproc_Presets.mrml|'''RegLib_C33_Preproc_Presets''' Presets for DWI resampling & DTI estimation <small> (.mrml file 8 kB) </small>]]''' | **[[Media:RegLib_C33_Preproc_Presets.mrml|'''RegLib_C33_Preproc_Presets''' Presets for DWI resampling & DTI estimation <small> (.mrml file 8 kB) </small>]]''' | ||
Revision as of 22:08, 23 November 2010
Home < Projects:RegistrationLibrary:RegLib C33Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
v3.6.1  Slicer Registration Library Exampe #33: Diffusion Weighted Image Volume: align with structural reference MRI
Slicer Registration Library Exampe #33: Diffusion Weighted Image Volume: align with structural reference MRI
Input
|
|
|
|
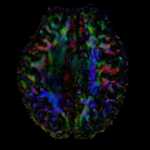
|
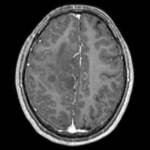
| fixed image/target T1 |
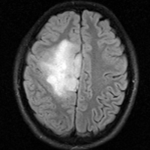
fixed image/target FLAIR |
moving image 2a DTI baseline |
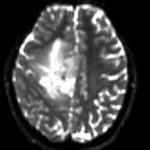
moving image 2b DTI tensor |
Modules
- Slicer 3.6.1 recommended modules: BrainsFit
Objective / Background
This is a typical example of DTI processing. Goal is to align the DTI image with a structural scan that provides accuracte anatomical reference. The DTI contains acquisition-related distortion and insufficient contrast to discern anatomical detail. For treatment planning and evaluation, location of functionally critical fiber tracts relative to the pathology is sought.
Keywords
MRI, brain, head, intra-subject, DTI, DWI
Input Data
- reference/fixed : T1 axial, 0.98 x 0.98 x 1 , 192 x 256 x 176
- reference/fixed : FLAIR axial, 0.4mm resolution in plane, 4mm slices, 448 x 512 x 30
- moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 1.96 x 1.96 x 3 mm voxel size, oblique, 128 x 128 x 40
- moving: Tensor data of DTI volume, oblique, same orientation as Baseline image. The result Xform will be applied to this volume. The original DWI has 64 directions, the extracted DTI volume has 9 scalars, i.e. 128 x 128 x 40 x 9
Overall Strategy
- Register FLAIR to T1 (affine) & resample
- Resample DWI to isotropic voxel size: use T1 as size reference
- Convert DWI -> DTI, incl. mask & baseline extraction
- Register DTI_baseline to FLAIR (affine+nonrigid) w/o masking
- Resample DTI with above transform
Procedures
- Phase I: Load & Convert DICOM
- run Dicom2NRRD converter for the DWI Dicom set. Output file: DWI.nrrd
- load DWI.nrrd
- load T1, FLAIR and T1Gd images via AddVolume...
- Subsample T1: The T1 is 512 x 512 x 176 (0.5 x 0.5 x 1) resolution, which is too large as a reference to sample the DTI into. So we generate a smaller, isotropic 1x1x1 version of the T1 that can serve as reference:
- Open Filtering/Resample Scalar Volume module
##Input: T1, Output: new volume, rename to T1_sub
- interpolation: Hamming
- Spacing: 1,1,1
- Phase II: Check initial Alignment and anisotropy
- large initial misalignment and high voxel anisotropy can cause problems when registering and resampling the DTI. In such cases it is advisable to resample the DWI first into a more isotropic space and, if necessary, initially realign to a similar orientation as the T1 reference, 'before converting to DTI. In this example DWI is sufficiently aligned and has voxel dimensions of 1 x 1 x 2.6, which is borderline. Anisotropy ratios greater than 1:3 tend to benefit from initial resampling. In this case we proceed without.
- 'Phase III: convert DWI -> DTI
- go to module: Diffusion / Utilities / Diffusion Tensor Estimation
- input: DWI, output : DTI, DTI_base and DTI_mask
- Input DWI Volume: DWI, Output DTI Volume: create new, rename to "DTI"
- Output Baseline: create new, rename to "DTI_base"
- Otsu Threshold Mask: create new, rename to "DTI_iso_mask"
- check boxes for Remove Islands and Apply Mask
- leave defaults
- Phase IVa: Register DTI-T1 (unmasked)
- open Registration : BrainsFit module (presets: Xf1_DTI-T1_unmasked
- Registration Phases:
- set T1_sub as fixed and DTI_base as moving image
- select/check Include Affine registration phase
- select/check Include BSpline registration phase
- Output Settings:
- select a new transform "Slicer BSpline Transform", rename to "Xf1_DTI-T1_unmasked"
- select a new volume "Output Image Volume, rename to "DTI_base_Xf1"
- Registration Parameters: increase Number Of Samples to 200,000
- Registration Parameters: set Number Of Grid Subdivisions to 5,5,3
- Leave all other settings at default
- click: Apply; (runtime < 1 min 10 sec. on MacPro Quad Core 2.4Ghz)
- result: the absence of the skull in the DTI causes significant and incorrect stretch of image in the I-S direction.
- we repeat with mask, using the above estimate to produce a mask for the T1
- Phase IVb: Build mask for T1: resample DTI_mask
- Filtering/Resample Scalar Vector DWI Volume
- Use provided presets or set the following:
- input: DTI_mask
- reference: T1_sub
- output: create new, rename to T1_sub_mask
- Transform node: Xf1_DTI-T1_unmasked
- Transforms order: check output-to-input
- interpolation type: check "nn"
- click: Apply
- Go to Volumes module
- select new "T1_sub_mask: volume, go to Info tab
- check the labelmap checkbox
- Phase IVc: Register DTI-T1 (masked)
- open Registration : BrainsFit module (presets: Xf1_DTI-T1_masked)
- settings as in Phase IVa above, except:
- select a new transform "Slicer BSpline Transform", rename to "Xf2_DTI-T1_masked"
- select a new volume "Output Image Volume, rename to "DTI_base_Xf2"
- Mask Processing Mode: check "ROI"
- Input Fixed Mask: T1_mask
- Input Moving Mask: DTI_mask
- Apply. Runtime 45 sec.
- go to Volumes module to adjust window&level for new result "DTI_base_Xf2"
- alignment is now much improved
- settings as in Phase IVa above, except:
- Phase V: Resample DTI
- Open the Resample DTI Volume module (found under: All Modules)
- Input Volume: select DTI
- Output Volume: select New DTI Volume, rename to DTI_Xf2
- Reference Volume: select T1_sub
- Transform Parameters: select transform "Xf2_DTI-T1_masked
- check box: output-to-input
- Leave all other settings at defaults
- Click Apply; runtime ~ 3 min.
- Go to the Volumes module, select the newly produced DTI_Xf2 volume
- under the Display tab, select Color Orientation from the Scalar Mode menu
- set T1 as background and new DTI_Xf2 volume as foreground
- Set fade slider to see DTI overlay onto the T2 image
for more details see the tutorial(s) under Downloads above
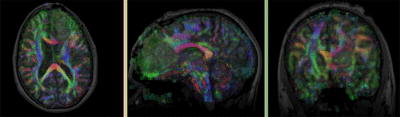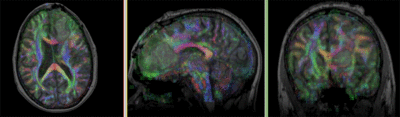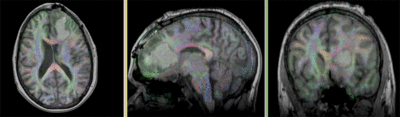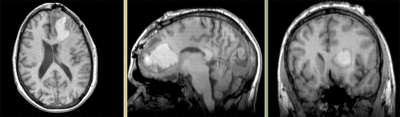
Registration Results
Download
- Data
- Presets
- Documentation
Discussion: Registration Challenges
- The DTI contains acquisition-related distortions (commonly EPI acquisitions) that can make automated registration difficult.