Projects:RegistrationLibrary:RegLib C11
From NAMIC Wiki
Home < Projects:RegistrationLibrary:RegLib C11
Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
Slicer Registration Library Exampe #11: Co-registration of two anatomic brain atlases for merging of structure labels
Objective / Background
This is an example of inter-subject registration via surface matching. The structures of interest are a small subset of the entire image, hence registration is not driven by image intensities but rather two model surfaces derived from the labelmaps.
Keywords
MRI, brain, head, inter-subject, atlas to atlas, surface-based segmentation, thalamic nuclei
Input Data
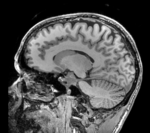
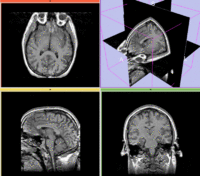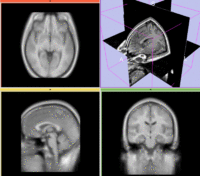
 reference/fixed : T1w coronal, 1mm isotropic. Called A1_gray
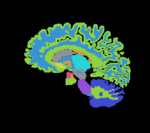
reference/fixed : T1w coronal, 1mm isotropic. Called A1_gray reference/fixed : labelmap , aligned with above. Called A1_label
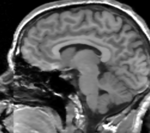
reference/fixed : labelmap , aligned with above. Called A1_label moving: T1w coronal, 0.9 inplane, 1.5mm coronal slices. Called A0_gray
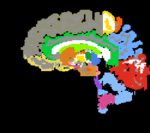
moving: T1w coronal, 0.9 inplane, 1.5mm coronal slices. Called A0_gray moving: labelmap , aligned with above. Called A0_label
moving: labelmap , aligned with above. Called A0_label
Methods
- Visualize & browse A0 data: determine label range of thalamic nuclei labels in A0_label: 500-526
- Visualize & browse A1 data: determine label range of thalamus lables in A1_label: 10 and 49
- Build label mask of thalamus for A0: threshold A0_label from 500 t0 526
- Build label mask of thalamus for A1: change label 10 to 49; threshold A1_label from 49 to 49
- Smooth A1 thalamus labelmap: Median Filter
- Build thalamus surface model of A0
- Build thalamus surface model of A1
- run Python Surface ICP Registration , Settings:
- Landmark transform mode: Affine
- Mean Distance Mode: RMS
- Maximum Number of Iterations: 500
- Maximum Nmber of Landmarks: 200
- Start by matching centroids: yes
- maximum mean distance: 0.01
- Initial Transform: none
- Input Surface: A0_ThalamusModel
- Target Surface: A1_ThalamusModel
- Output Surface: none
- Output Transform: Xform_A0Affine_ICP
- Apply Registration Xform to labelmap: Resample Scalar/Vector/DWI Volume module. Settings:
- Input Volume : A0_labels
- Reference Volume : A1_labels
- Output Volume : “Create New Volume”, rename to “A0_labels_aff”
- Transform Node: Xform_A0Affine_ICP
- Interpolation Type: “nn”
- Change datatype of new volume A0_labels_aff to labelmap (Volumes module)
- Apply Registration Xform to thalamus mask: Resample Scalar/Vector/DWI Volume module. Settings:
- Input Volume : A0_thalamus
- Reference Volume : A1_labels
- Output Volume : “Create New Volume”, rename to A0_thalamus_aff
- Transform Node: Xform_A0Affine_ICP
- Interpolation Type: “nn”
- Change datatype of new volume A0_labels_aff to labelmap (Volumes module)
- Mask new labelmap with A0_thalamus_aff: Mask Image module
- Input Volume : A0_labels_aff
- Mask Volume : A0_thalamus_aff�
- Masked Volume: A0_labels aff
- Mask new labelmap with A1_thalamus: Mask Image module
- Input Volume : A0_labels_aff�
- Mask Volume : A1_thalamus_aff�
- Masked Volume: New volume, rename to: A0_labels aff_clip
- Typecast A1_labelmap: Cast Image module
- Input Volume: A1_label
- Output Volume: A1_label
- Output Type: “short”
- Merge Labelmaps:Image Label Combine module
- Input Label Map A: A0_label_aff_clip
- Input Label Map B: A1_labels
- Output Label Map: “Create New Volume”, rename to A1_labels_merged
- First label overwrites second: yes
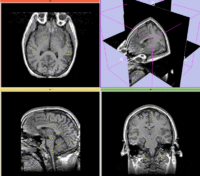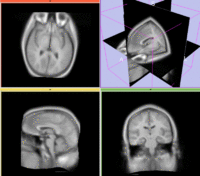
Registration Results
Download
- download entire package (Data,Tutorial, Solution, zip file 14 MB)
- Presets
- Tutorial only
- Image Data only
Discussion: Registration Challenges
- Because the atlas represents a probabilistic image (i.e. contains blurring from combining multiple subjects), its contrast differs significantly from the target image.
- The atlas has strong rotational misalignment that can cause difficulty for automated affine registration.
- The two images represent different anatomies, a non-rigid registration is required
Discussion: Key Strategies
- Because of the strong differences in image contrast, Mutual Information is recommended as the most robust metric.
- masking (skull stripping) is highly recommended to obtain good results.
- because speed is not that critical, we increase the sampling rate for both affine and BSpline registration
- we also expect larger differences in scale & distortion than with regular structural scans: so we significantly (2x-3x) increase the expected values for scale and skew from the defaults.
- a good affine alignment is important before proceeding to non-rigid alignment to further correct for distortions.
Acknowledgments
- dataset provided by Killian Pohl, Ph.D.