Difference between revisions of "DBP2:Harvard:Brain Segmentation Roadmap"
| (247 intermediate revisions by 8 users not shown) | |||
| Line 1: | Line 1: | ||
| + | Back to [[NA-MIC_Internal_Collaborations|NA-MIC Collaborations]], [[DBP2:Harvard|Harvard DBP 2]] | ||
| + | __NOTOC__ | ||
| + | =Stochastic Tractography for VCFS= | ||
== Roadmap == | == Roadmap == | ||
| − | + | The main goal of this project is to develop end-to-end application that would be used to characterize anatomical connectivity abnormalities in the brain of patients with velocardiofacial syndrome (VCFS), and to link this information with deficits in schizophrenia. This page describes the technology roadmap for stochastic tractography, using newly acquired 3T data, NAMIC tools and slicer 3. | |
| − | |||
| − | + | == Algorithm == | |
| − | ; A - | + | [[Image:IC_sto_new.png|thumb|right|200px|<font size=1> Figure 1: Comparison of deterministic and stochastic tractography algorithms</font>]] |
| − | + | ; A-Description | |
| − | + | * Most tractography methods estimate fibers by tracing the maximum direction of diffusion. A limitation of this approach is that, in practice, several factors introduce uncertainty in the tracking procedure, including, noise, splitting and crossing fibers, head motion and image artifacts. To address this uncertainty, stochastic tractography methods have been developed to quantify the uncertainty associated with estimated fibers (Bjornemo et al., 2002). Method uses a propagation model based on stochastics and regularization, which allows paths originating at one point to branch and return a probability distribution of possible paths. The method utilizes principles of a statistical Monte Carlo method called Sequential Importance Sampling and Resampling (SISR). Based on probability functions, using a sequential importance sampling technique ([http://lmi.bwh.harvard.edu/papers/pdfs/2002/bjornemoMICCAI02.pdf Bjornemo et al., 2002]), one can generate thousands of fibers starting in the same point by sequentially drawing random step directions. This gives a very rich model of the fiber distribution, as contrasted with single fibers produced by conventional tractography methods. Moreover, from a large number of sampled paths, probability maps can be generated, providing better estimates of connectivity between several anatomical locations. A comparison of the algorithms can be seen here. (Figure 1) | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| + | [[Image:StochasticPic.PNG|thumb|right|200px|<font size=1>Figure 2: Stochastic tractography of uncinate fasciculis on anatomical data (left) and cingulum bungle on fMRI scan (right)</font>]] | ||
| + | ; B-Possible Applications | ||
| + | * Since diffusion direction uncertainty within the gray matter is quite significant; principal diffusion direction approaches usually do not work for tracking between two gray matter regions. Thus if one requires finding connections between a priori selected anatomical gray matter regions, defined either by anatomical segmentations (in case of using structural ROI data), or functional activations (in case of megring DTI with fMRI), stochastic tractography seems to be the method of choice. Here is an example of this application to anatomical data (Figure 2, left image) and to fMRI data (Figure 2, right image). | ||
| − | === | + | * Stochastic Tractography is also comparable, if not better, in defining large white matter fiber bundles, especially those traveling through white matter regions characterized by increased diffusion uncertainty (fiber crossings). Example of such application to internal capsule. (Figure 3) |
| − | * | + | |
| − | * | + | [[Image:IC-comp-new.png|thumb|right|200px|<font size=1>Figure 3: Streamline vs. stochastic tractography of the Internal Capsule</font>]] |
| + | ; C-References | ||
| + | |||
| + | * [http://lmi.bwh.harvard.edu/papers/pdfs/2002/bjornemoMICCAI02.pdf Björnemo M, Brun A, Kikinis R, Westin CF. Regularized stochastic white matter tractography using diffusion tensor MRI. In Fifth International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI'02). Tokyo, Japan, 2002;435-442.] | ||
| + | * [http://lmi.bwh.harvard.edu/papers/pdfs/2006/frimanTMI06.pdf Friman, O., Farneback, G., Westin CF. A Bayesian Approach for Stochastic White Matter Tractography. IEEE Transactions on Medical Imaging, Vol 25, No. 8, Aug. 2006] | ||
| + | |||
| + | [[Image:StochasticGUI1.PNG|thumb|right|200px|<font size=1> Figure 4: Python Stochastic Tractography GUI </font>]] | ||
| + | |||
| + | ==Module== | ||
| + | Can be found in: MODULES > PYTHON MODULES > PYTHON STOCHASTIC TRACTOGRAPHY | ||
| + | ;Functionality of Python Stochastic Tractography module in Slicer 3.0 | ||
| + | * IO: | ||
| + | Module reads files (DWI and ROIs) in nhdr format. | ||
| + | * Smoothing: | ||
| + | One can smooth the DWI data (only Gausian smoothing is supported at this time). We recommend it if the data is noisy. | ||
| + | * Brain Mask: | ||
| + | The Brain mask defines the volume in which the tensor will be computed and the tracts evaluated. If Enabled, will use threshold values on the baseline instead of WM Mask defined in IO panel. | ||
| + | *IJK/RAS Switch | ||
| + | Chooses the way the nhdrs are read. | ||
| + | *Diffusion Tensor: | ||
| + | This step allows output of the tensor image and can output anisotropy indices (FA/Mode/Trace) | ||
| + | *Tractography: | ||
| + | Parameters that need to be adjusted: | ||
| + | :1. Total Tracts: The amount of tracts that will be seeded from each voxel (we recommend between 500 and 1000 tracts, depending on the workstation power- 1000 tracts per voxel seeded within the large ROI for high resolution DWI can take a long time to compute). | ||
| + | :2. Maximum tract length: (in mm) This can eliminate long, unwanted tracts if the regions for which connection is measured are located close to each other | ||
| + | :3. Step Size(mm): distance between each re-estimation of tensors, usually between 0.5 and 1 mm. Adjust to make sure step size is not larger than the voxel spacing in any direction, which would allow voxels to be "jumped over." | ||
| + | :4. Stopping criteria: This can be used on the top of WM mask to terminate tracts when FA drops below supplied threshold (in case they frequently travel through CSF, for example). | ||
| + | :5. Use Basic Method: switches between Friman and McGraw algorithms. | ||
| + | *Connectivity Map: | ||
| + | This step creates output probability maps. | ||
| + | :1. binary: each voxel is counted only once if at least one fiber pass through it | ||
| + | :2. cumulative: tracts are summed by voxel independently | ||
| + | :3. weighted: tracts are summed by voxel depending on their length | ||
| + | *Length Based: | ||
| + | This step will output only either the shortest 1/3, middle 1/3, or longest 1/3 of the tracts. | ||
| + | *Threshold | ||
| + | This step will reject tracts whose endpoints are lower than the threshold value. | ||
| + | *Spherical ROI vicinity | ||
| + | This will make the ROI a sphere based on the ROI’s center of gravity (with the sphere’s radius being the distance from the center to the ROI’s furthest point). This sphere can be inflated by raising the Vicinity level to the number of steps you’d like to increase the ROI’s size by. | ||
| + | *Vicinity | ||
| + | This step traces back n number of steps from tract endpoint to check if track crosses target ROI. If so, tract is included. | ||
| + | |||
| + | Then, probability maps can be saved as ROIs, and either used directly, or thresholded (at certain probability, step claimed by few publications to remove noise) in slicer to mask and compute average FA, Mode, Trace for entire connection. Diffusion indices can be also weighted by the probability of connection for each voxel. | ||
| + | |||
| + | :* Module documentation and training data can be found here: | ||
| + | :**[[Slicer3:Stochastic_tractography]] | ||
| + | |||
| + | ==Work Accomplished== | ||
| + | [[Image:helix_withsmoothing.png|thumb|right|200px|<font size=1>Figure 6: Stochastic tractography from a single ROI on helix phantom</font>]] | ||
| + | ; A - Optimization and testing of stochastic tractography algorithm : | ||
| + | :* Original methodological paper, as well as our first attempts to use the algorithm (CC+ and matlab scripts) have been done on old "NAMIC" 1.5T LSDI data ([https://portal.nbirn.net:443/gridsphere/gridsphere?cid=srbfilebrowser&gs_action=gotoDirectory&CPq_dirpath=%2Fhome%2FProjects%2FNAMIC__0003%2FFiles%2FHarvard%2Farchive_morph&up=CPq&JavaScript=enabled Structural MRI] and [https://portal.nbirn.net:443/gridsphere/gridsphere?cid=srbfilebrowser&gs_action=gotoDirectory&up=7li&7li_dirpath=%2Fhome%2FProjects%2FNAMIC__0003%2FFiles%2FHarvard%2Farchive_diffusion&JavaScript=enabled DTI data]). | ||
| + | :* Algorithm has been optimized to work on higher resolution data (available here: [https://portal.nbirn.net/gridsphere/gridsphere?cid=srbfilebrowser&gs_action=moveUpDir&gv7_dirpath=%2Fhome%2FProjects%2FNAMIC__0003%2FFiles%2FPNL%2F3T_strct_dti_fmri%2Fcase01017&up=gv7&JavaScript=enabled|PNL 3T Data]). | ||
| + | :* Tests have been done also on the spiral diffusion phantom, to make sure diffusion directions and scanner coordinates are handled properly by the algorithm (Figure 6). | ||
| + | :* Separate components of work flow have been tested, including the masks, and impact of their precision on stochastic output, as well as impact of number of seeding points on tractography results. | ||
| + | :* Module have been tested on other Philips and GE datasets. | ||
| + | |||
| + | ; B - Clinical Applications | ||
| + | |||
| + | :* Algorithm was used to trace and analyze anterior limb of the internal capsule on 1.5T data. It generated reacher representation of frontal fiber projections, it also turned out to be more sensitive to group differences in white matter integrity that conventional deterministic tractography (see Figure 3). | ||
| + | :* Algorithm was also used to trace smaller white matter fiber tracts, such as Cingulum, Fornix, Uncinate Fasciculus, Arcuate Fasciculus on 3T "Santa Fe" dataset (http://www.na-mic.org/Wiki/index.php/SanteFe.Tractography.Conference) | ||
| + | |||
| + | == Related Clinical Projects == | ||
| + | |||
| + | ;* Arcuate Fasciculus Extraction and Analysis in Schizophrenia | ||
| + | [[Image:STArcuate.jpg|thumb|right|200px|<font size=1>Figure 7: The arcuate fasciculus including seed, midpoint and target ROI's.</font>]] | ||
| + | We are investigating Arcuate Fasciculus using Stochastic Tractography (Figure 7). This structure is especially important in both VCFS and schizophrenia, as it connects language related areas (Brocka and Wernicke's), and is involved in language processing quite disturbed in schizophrenia patients. It also can not be reliably traced using deterministic tractography. | ||
| + | :Project involves: | ||
| + | :* Whole brain segmentation, and automatic extraction of regions interconnected by Arcuate Fasciculus (Inferior frontal and Superior Temporal Gyri). | ||
| + | :* White matter segmentation, in order to prevent algorithm from traveling through the ventricles, where diffusivity is high. | ||
| + | :* Non-linear registration of labelmaps to the DTI space. | ||
| + | :* Seeding tracts within both ROIs. | ||
| + | :* Extracting path of interest, and calculating FA along the path for group comparison. Presentation of previous results for 7 schizophrenics and 12 control subjects, can be found here: [[Media:NAMIC_AHM_Arcuate.ppt|Progress Report Presentation]]. | ||
| + | :* Results presentation and paper submission. Abstract was accepted and results presented at the World Biological Psychiatry Symposium in Venice, Italy. Paper has been submitted to Neuroimage. | ||
| + | |||
| + | [[Image:connectivity1.jpg|thumb|right|200px|<font size=1>Figure 8: Maps of inferior frontal cortex (Broca) connectivity in controls and patients with schizophrenia.</font>]] | ||
| + | |||
| + | ;* Semantic Network Connectivity in Schizophrenia | ||
| + | We use combination of fMRI and DTI data to define and characterize functional and anatomical connectivity within the semantic processing network in schizophrenia. | ||
| + | :Project involves: | ||
| + | :* fMRI data analysis and identification of functional nodes involved in semantic processing in healthy controls and seubjects with schizophrenia | ||
| + | :* Analysis of functional connectivity (using FSL) between nodes of semantic network | ||
| + | :* Whole brain Voxel Based analysis of DTI data in same population | ||
| + | :* Use of stochastic tractography to identify connections between functional nodes | ||
| + | :* Correlational analysis involving anatomical and functional connectivity data. | ||
| + | :* Results presentation and paper submission. Data was presented at HBM conference in 2007, paper has been accepted for publication at HBM. | ||
| + | :* <font color="red">'''New: '''</font>June 10th 2009, paper accepted for publication in Human Brain Mapping: "Functional and Anatomical Connectivity Abnormalities in Left Inferior Frontal Gyrus in Schizophrenia" by Jeong, Wible, Hashimoto and Kubicki, HBM in Press | ||
| + | |||
| + | [[Image:ScreenshotFreeSurferDeepMatterSagitalView-vcase1-2009-06-12.jpg|thumb|right|200px|<font size=1>Figure 9: Brain automatic segmentation of subject with VCFS.</font>]] | ||
| + | |||
| + | ;* Anatomical Connectivity Abnormalities in VCFS | ||
| + | We use combination of structural MRI and DTI to investigate anatomical abnormalities that would characterize patients with VCFS, and relationship of these abnormalities to those observed in schizophrenia. | ||
| + | :Project involves: | ||
| + | :* Whole brain, automatic segmentation of brain images obtained from patients with VCFS in order to identify structures involved in this disease. | ||
| + | :* Registration between anatomical and DTI scans (manual skull stripping followed by linear followed by nonlinear registration of SPGRs to DTI space). | ||
| + | :* Use of stochastic tractography to identify connections between gray matter regions identified in the disease. | ||
| + | :* Extracting paths of interest, and calculating FA along the paths for group comparison. | ||
| + | :* Correlational analysis involving anatomical and connectivity data, clinical information and genetic data. | ||
| + | :* Results presentation and paper submission. We plan to submit an abstract with study results for Biological Psychiatry symposium (dedline December 2010). | ||
| + | |||
| + | [[Image:Anna.png|thumb|right|200px|<font size=1>Figure 10: Max Plank data showing tracts through the corpus connecting 2 cortical ROIs defined by fMRI activations.</font>]] | ||
| + | |||
| + | ;* Study of Default Network in Psychosis | ||
| + | We are collaborating with the Department of Neurophysiology Max Planck Institute in Frankfurt (Anna Rotarska contact person), where we use stochastic tractography to measure integrity of anatomical connections within the default network in schizophrenia. The fMRI resting state results (ROIs) have been co-registered with anatomical scans, and then again registered to DTI space. Figure 8 shows pilot data of the white matter connections through the corpus callosum between the left and the right auditory cortex ROIs as defined by fMRI data. | ||
| + | :* Results presentation and paper submission. We are in the process of analyzing results. | ||
| + | |||
| + | ;* Study of OFC-ACC connectivity in Chronic Schizophrenia | ||
| + | We are using stochastic tractography to examine white matter connectivity between medial anterior and posterior OFC and rostral ACC (cognitive part of ACC). | ||
| + | :* Results presentation and paper submission. Data was analyzed for 27 SZ and 256 HC subjects. Schizophrenia group demonstrated significant mean FA reduction in the connection between left anterior OFC and ACC, and between bilateral posterior OFC. Poster was presented at Annual Biological Psychiatry Meeting, paper in preparation. | ||
| + | |||
| + | [[Image:Otani-NAMIC.png|thumb|right|200px|<font size=1>Figure 11: Stochastic density map (green) between OFC (red )and ACC (purple).</font>]] | ||
| + | |||
| + | ;* Study of thalamus segmentation based on cortical connectivity in Chronic Schizophrenia | ||
| + | We use stochastic tractography to segment thalamus into discrete ROIs based on its connectivity to 10 cortical ROIs (for each hemisphere) of the frontal lobe. Cortical ROIs are extracted using free surfer, and co-registered to DTI space using FSL. Thalamus is painted by connectivity using additional in-house matlab script (available after request). Volumes of the thalamic ROIs as well as FA for each individual connection are our measures of interest. | ||
| + | :* Results presentation and paper submission. Data analysis and paper in preparation. | ||
| + | |||
| + | [[Image:Taiga-NAMIC.png|thumb|right|400px|<font size=1>Figure 12: Thalamus parcellation into discrete ROIs based on its connectivity to 10 cortical ROIs using stochastic tracking.</font>]] | ||
| + | |||
| + | ;* Tractography Comparison Project | ||
| + | We are also working on a [http://www.na-mic.org/Wiki/index.php/SanteFe.Tractography.Conference tractography comparison project]dataset, where we apply stochastic tractography to phantom, as well as test dataset. | ||
| + | |||
| + | ; - Related References | ||
| + | |||
| + | :* Shenton, M.E., Ngo, T., Rosenberger, G., Westin, C.F., Levitt, J.J., McCarley, R.W., Kubicki, M. Study of Thalamo-Cortical White Matter Fiber Tract Projections in Schizophrenia Using Diffusion Stochastic Tractography. Poster presented at the 46th Meeting of the American College of Neuropsychopharmacology, Boca Raton, FL, December 2007. | ||
| + | :* Terry DP, Rausch AC, Alvarado JL, Melonakos ED, Markant D, Westin CF, Kikinis R, de Siebenthal J, Shenton ME, Kubicki M. White Matter Properties of Emotion Related Connections in Schizophrenia. Poster presented at the 2009 Mysell Poster Day, Dept. of Psychiatry, Harvard Medical School, April 2009] | ||
| + | :* Jorge_poster.pdf Alvarado JL, Terry DP, Markant D, Ngo T, Kikinis R, Westin CF, McCarley RW, Shenton ME, Kubicki M. Study of Language-Related White Matter Tract Connections in Schizophrenia using Diffusion Stochastic Tractography. Poster presented at the 2009 Mysell Poster Day, Dept. of Psychiatry, Harvard Medical School, April 2009] | ||
| + | :* Melonakos ED, Shenton ME, Markant D, Alvarado J, Westin CF, Kubicki M. White Matter Properties of Orbitofrontal Connections in Schizophrenia. Poster being presented at the 64th Meeting of the Society of Biological Psychiatry. Vancouver, BC. May 2009. | ||
| + | :* Kubicki, M. Khan, U., Bobrow, L., O'Donnell, L. Pieper, S. Westin, CF., Shenton, ME. New Methods for Assessing Whole Brain DTI Abnormalities in Schizophrenia. Presentation given at the International Congress of World Psychiatric Association. Florence, Italy. April 2009. | ||
| + | :* Kubicki, M., Markant, D., Ngo, T., Westin, CF., McCarley, RW., Shenton, ME. Study of Language Related White Matter Fiber Tract Projections in Schizophrenia Using Diffusion Stochastic Tractography. Presentation given at the International Congress of World Psychiatric Association. Florence, Italy. April 2009. | ||
| + | :* T. Otani, M Kubicki, S. Bouix, P Nestor, A Rausch, T Asami, D, Terry, E Melonakos, K Hawley, P Pelavin, J Alvarado, A LaVenture, J Siebenthal, R McCarley, and M Shenton. White matter connections between orbitofrontal cortex and anterior cingulate cortex in shchizophrenia, Annual Meeting, Society of Biological Psychiatry, 2010. | ||
===Staffing Plan=== | ===Staffing Plan=== | ||
| − | * Sylvain and | + | * Sylvain and Ryan are the DBP resources charged with adapting the tools in the NA-MIC Kit to the DBP needs |
| + | * Andrew Rauch is our NAMIC RA. | ||
| + | * Ryan is our NAMIC software engineer. He is responsible for improving the STM (Stochastic Tractography Module), and making sure software works with STM compliant datasets. | ||
* Polina is the algorithm core contact | * Polina is the algorithm core contact | ||
| − | * Brad is the engineering core contact | + | * Brad is the engineering core contact |
===Schedule=== | ===Schedule=== | ||
| − | * ''' | + | * '''10/2007''' - Optimization of Stochastic Tractography algorythm for 1.5T data. |
| − | * '''12/ | + | * '''10/2007''' - Algorythm testing on Santa Fe data set and diffusion phantom. |
| − | * ''' | + | * '''06/2008''' - Optimization of Stochastic Tractography algorythm for 3T data. |
| − | * '''01/ | + | * '''11/2008''' - Slicer 3 module prototype using python. |
| − | * ''' | + | * '''12/2008''' - Slicer 3 module official release |
| − | * '''07/ | + | * '''12/2008''' - Documentation and packaging for dissemination. |
| − | * ''' | + | * '''12/2008''' - First clinical application of stochastic tractography module. |
| − | * ''' | + | * '''01/2009''' - First draft of the clinical paper. |
| − | * '''01/ | + | * '''05/2009''' - Distortion correction and nonlinear registration added to the module |
| + | * '''05/2009''' - Symposium on tractography, including stochastic methods at World Biological Psychiatry Symposium in Florence, Italy. | ||
| + | * '''05/2009''' - Presentation of Arcuate Fasciculus findings at World Biological Psychiatry Symposium in Florence, Italy. | ||
| + | * '''07/2009''' - Summer Programming week- work on optimizing and speeding up data processing, releasing second generation of software that includes preprocessing pipeline. | ||
| + | * '''07/2009''' - Continue working on clinical collaborative studies using stochastic tractography module. | ||
| + | * '''12/2009''' - Submission of abstracts reporting findings of several clinical studies involving stochastic tractography, including anatomical connectivity abnormalities in patients with VCFS and anatomical and functional connectivity abnormalities in schizophrenia. | ||
| + | * '''01/2010''' - AHM progress presentation. | ||
| + | * '''05/2010''' - Presentation of clinical findings at Annual Biological Psychiatry Symposium, San Francisco. | ||
| + | * '''07/2010''' - Summer Programming week- presentation of new software manual, software included in new slicer 3 release. | ||
===Team and Institute=== | ===Team and Institute=== | ||
*PI: Marek Kubicki (kubicki at bwh.harvard.edu) | *PI: Marek Kubicki (kubicki at bwh.harvard.edu) | ||
| − | *DBP2 Investigators: Sylvain Bouix, | + | *DBP2 Investigators: Sylvain Bouix, Yogesh Rathi, Ryan Ecbo |
*NA-MIC Engineering Contact: Brad Davis, Kitware | *NA-MIC Engineering Contact: Brad Davis, Kitware | ||
*NA-MIC Algorithms Contact: Polina Gollard, MIT | *NA-MIC Algorithms Contact: Polina Gollard, MIT | ||
| − | === | + | ===Publications=== |
| + | |||
| + | ''In print'' | ||
| + | |||
| + | *[http://www.na-mic.org/pages/Special:Publications?text=Kubicki+AND+Westin+AND+DTI&submit=Search&words=all&title=checked&keywords=checked&authors=checked&abstract=checked&sponsors=checked&searchbytag=checked| NA-MIC Publications Database - Clinical Applications] | ||
| + | |||
| + | *[http://www.na-mic.org/pages/Special:Publications?text=Ngo+AND+Golland&submit=Search&words=all&title=checked&keywords=checked&authors=checked&abstract=checked&sponsors=checked&searchbytag=checked| NA-MIC Publications Database - Algorithms Development] | ||
| + | |||
| + | |||
| + | [[Category: Schizophrenia]] [[Category: Diffusion MRI]] [[Category: Segmentation]] | ||
Latest revision as of 20:24, 2 November 2010
Home < DBP2:Harvard:Brain Segmentation RoadmapBack to NA-MIC Collaborations, Harvard DBP 2
Stochastic Tractography for VCFS
Roadmap
The main goal of this project is to develop end-to-end application that would be used to characterize anatomical connectivity abnormalities in the brain of patients with velocardiofacial syndrome (VCFS), and to link this information with deficits in schizophrenia. This page describes the technology roadmap for stochastic tractography, using newly acquired 3T data, NAMIC tools and slicer 3.
Algorithm
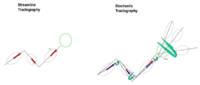
- A-Description
- Most tractography methods estimate fibers by tracing the maximum direction of diffusion. A limitation of this approach is that, in practice, several factors introduce uncertainty in the tracking procedure, including, noise, splitting and crossing fibers, head motion and image artifacts. To address this uncertainty, stochastic tractography methods have been developed to quantify the uncertainty associated with estimated fibers (Bjornemo et al., 2002). Method uses a propagation model based on stochastics and regularization, which allows paths originating at one point to branch and return a probability distribution of possible paths. The method utilizes principles of a statistical Monte Carlo method called Sequential Importance Sampling and Resampling (SISR). Based on probability functions, using a sequential importance sampling technique (Bjornemo et al., 2002), one can generate thousands of fibers starting in the same point by sequentially drawing random step directions. This gives a very rich model of the fiber distribution, as contrasted with single fibers produced by conventional tractography methods. Moreover, from a large number of sampled paths, probability maps can be generated, providing better estimates of connectivity between several anatomical locations. A comparison of the algorithms can be seen here. (Figure 1)
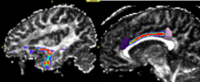
- B-Possible Applications
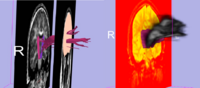
- Since diffusion direction uncertainty within the gray matter is quite significant; principal diffusion direction approaches usually do not work for tracking between two gray matter regions. Thus if one requires finding connections between a priori selected anatomical gray matter regions, defined either by anatomical segmentations (in case of using structural ROI data), or functional activations (in case of megring DTI with fMRI), stochastic tractography seems to be the method of choice. Here is an example of this application to anatomical data (Figure 2, left image) and to fMRI data (Figure 2, right image).
- Stochastic Tractography is also comparable, if not better, in defining large white matter fiber bundles, especially those traveling through white matter regions characterized by increased diffusion uncertainty (fiber crossings). Example of such application to internal capsule. (Figure 3)
- C-References
- Björnemo M, Brun A, Kikinis R, Westin CF. Regularized stochastic white matter tractography using diffusion tensor MRI. In Fifth International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI'02). Tokyo, Japan, 2002;435-442.
- Friman, O., Farneback, G., Westin CF. A Bayesian Approach for Stochastic White Matter Tractography. IEEE Transactions on Medical Imaging, Vol 25, No. 8, Aug. 2006
Module
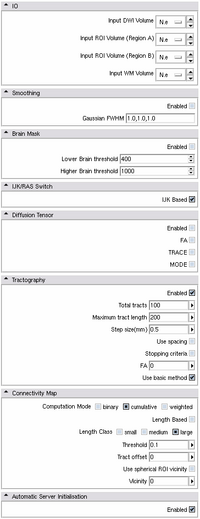
Can be found in: MODULES > PYTHON MODULES > PYTHON STOCHASTIC TRACTOGRAPHY
- Functionality of Python Stochastic Tractography module in Slicer 3.0
- IO:
Module reads files (DWI and ROIs) in nhdr format.
- Smoothing:
One can smooth the DWI data (only Gausian smoothing is supported at this time). We recommend it if the data is noisy.
- Brain Mask:
The Brain mask defines the volume in which the tensor will be computed and the tracts evaluated. If Enabled, will use threshold values on the baseline instead of WM Mask defined in IO panel.
- IJK/RAS Switch
Chooses the way the nhdrs are read.
- Diffusion Tensor:
This step allows output of the tensor image and can output anisotropy indices (FA/Mode/Trace)
- Tractography:
Parameters that need to be adjusted:
- 1. Total Tracts: The amount of tracts that will be seeded from each voxel (we recommend between 500 and 1000 tracts, depending on the workstation power- 1000 tracts per voxel seeded within the large ROI for high resolution DWI can take a long time to compute).
- 2. Maximum tract length: (in mm) This can eliminate long, unwanted tracts if the regions for which connection is measured are located close to each other
- 3. Step Size(mm): distance between each re-estimation of tensors, usually between 0.5 and 1 mm. Adjust to make sure step size is not larger than the voxel spacing in any direction, which would allow voxels to be "jumped over."
- 4. Stopping criteria: This can be used on the top of WM mask to terminate tracts when FA drops below supplied threshold (in case they frequently travel through CSF, for example).
- 5. Use Basic Method: switches between Friman and McGraw algorithms.
- Connectivity Map:
This step creates output probability maps.
- 1. binary: each voxel is counted only once if at least one fiber pass through it
- 2. cumulative: tracts are summed by voxel independently
- 3. weighted: tracts are summed by voxel depending on their length
- Length Based:
This step will output only either the shortest 1/3, middle 1/3, or longest 1/3 of the tracts.
- Threshold
This step will reject tracts whose endpoints are lower than the threshold value.
- Spherical ROI vicinity
This will make the ROI a sphere based on the ROI’s center of gravity (with the sphere’s radius being the distance from the center to the ROI’s furthest point). This sphere can be inflated by raising the Vicinity level to the number of steps you’d like to increase the ROI’s size by.
- Vicinity
This step traces back n number of steps from tract endpoint to check if track crosses target ROI. If so, tract is included.
Then, probability maps can be saved as ROIs, and either used directly, or thresholded (at certain probability, step claimed by few publications to remove noise) in slicer to mask and compute average FA, Mode, Trace for entire connection. Diffusion indices can be also weighted by the probability of connection for each voxel.
- Module documentation and training data can be found here:
Work Accomplished
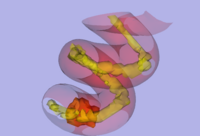
- A - Optimization and testing of stochastic tractography algorithm
-
- Original methodological paper, as well as our first attempts to use the algorithm (CC+ and matlab scripts) have been done on old "NAMIC" 1.5T LSDI data (Structural MRI and DTI data).
- Algorithm has been optimized to work on higher resolution data (available here: 3T Data).
- Tests have been done also on the spiral diffusion phantom, to make sure diffusion directions and scanner coordinates are handled properly by the algorithm (Figure 6).
- Separate components of work flow have been tested, including the masks, and impact of their precision on stochastic output, as well as impact of number of seeding points on tractography results.
- Module have been tested on other Philips and GE datasets.
- B - Clinical Applications
- Algorithm was used to trace and analyze anterior limb of the internal capsule on 1.5T data. It generated reacher representation of frontal fiber projections, it also turned out to be more sensitive to group differences in white matter integrity that conventional deterministic tractography (see Figure 3).
- Algorithm was also used to trace smaller white matter fiber tracts, such as Cingulum, Fornix, Uncinate Fasciculus, Arcuate Fasciculus on 3T "Santa Fe" dataset (http://www.na-mic.org/Wiki/index.php/SanteFe.Tractography.Conference)
Related Clinical Projects
- Arcuate Fasciculus Extraction and Analysis in Schizophrenia
We are investigating Arcuate Fasciculus using Stochastic Tractography (Figure 7). This structure is especially important in both VCFS and schizophrenia, as it connects language related areas (Brocka and Wernicke's), and is involved in language processing quite disturbed in schizophrenia patients. It also can not be reliably traced using deterministic tractography.
- Project involves:
- Whole brain segmentation, and automatic extraction of regions interconnected by Arcuate Fasciculus (Inferior frontal and Superior Temporal Gyri).
- White matter segmentation, in order to prevent algorithm from traveling through the ventricles, where diffusivity is high.
- Non-linear registration of labelmaps to the DTI space.
- Seeding tracts within both ROIs.
- Extracting path of interest, and calculating FA along the path for group comparison. Presentation of previous results for 7 schizophrenics and 12 control subjects, can be found here: Progress Report Presentation.
- Results presentation and paper submission. Abstract was accepted and results presented at the World Biological Psychiatry Symposium in Venice, Italy. Paper has been submitted to Neuroimage.
- Semantic Network Connectivity in Schizophrenia
We use combination of fMRI and DTI data to define and characterize functional and anatomical connectivity within the semantic processing network in schizophrenia.
- Project involves:
- fMRI data analysis and identification of functional nodes involved in semantic processing in healthy controls and seubjects with schizophrenia
- Analysis of functional connectivity (using FSL) between nodes of semantic network
- Whole brain Voxel Based analysis of DTI data in same population
- Use of stochastic tractography to identify connections between functional nodes
- Correlational analysis involving anatomical and functional connectivity data.
- Results presentation and paper submission. Data was presented at HBM conference in 2007, paper has been accepted for publication at HBM.
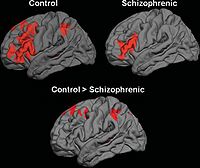
- New: June 10th 2009, paper accepted for publication in Human Brain Mapping: "Functional and Anatomical Connectivity Abnormalities in Left Inferior Frontal Gyrus in Schizophrenia" by Jeong, Wible, Hashimoto and Kubicki, HBM in Press
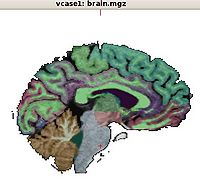
- Anatomical Connectivity Abnormalities in VCFS
We use combination of structural MRI and DTI to investigate anatomical abnormalities that would characterize patients with VCFS, and relationship of these abnormalities to those observed in schizophrenia.
- Project involves:
- Whole brain, automatic segmentation of brain images obtained from patients with VCFS in order to identify structures involved in this disease.
- Registration between anatomical and DTI scans (manual skull stripping followed by linear followed by nonlinear registration of SPGRs to DTI space).
- Use of stochastic tractography to identify connections between gray matter regions identified in the disease.
- Extracting paths of interest, and calculating FA along the paths for group comparison.
- Correlational analysis involving anatomical and connectivity data, clinical information and genetic data.
- Results presentation and paper submission. We plan to submit an abstract with study results for Biological Psychiatry symposium (dedline December 2010).
- Study of Default Network in Psychosis
We are collaborating with the Department of Neurophysiology Max Planck Institute in Frankfurt (Anna Rotarska contact person), where we use stochastic tractography to measure integrity of anatomical connections within the default network in schizophrenia. The fMRI resting state results (ROIs) have been co-registered with anatomical scans, and then again registered to DTI space. Figure 8 shows pilot data of the white matter connections through the corpus callosum between the left and the right auditory cortex ROIs as defined by fMRI data.
- Results presentation and paper submission. We are in the process of analyzing results.
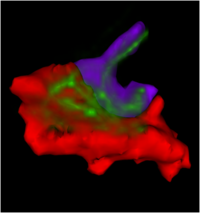
- Study of OFC-ACC connectivity in Chronic Schizophrenia
We are using stochastic tractography to examine white matter connectivity between medial anterior and posterior OFC and rostral ACC (cognitive part of ACC).
- Results presentation and paper submission. Data was analyzed for 27 SZ and 256 HC subjects. Schizophrenia group demonstrated significant mean FA reduction in the connection between left anterior OFC and ACC, and between bilateral posterior OFC. Poster was presented at Annual Biological Psychiatry Meeting, paper in preparation.
- Study of thalamus segmentation based on cortical connectivity in Chronic Schizophrenia
We use stochastic tractography to segment thalamus into discrete ROIs based on its connectivity to 10 cortical ROIs (for each hemisphere) of the frontal lobe. Cortical ROIs are extracted using free surfer, and co-registered to DTI space using FSL. Thalamus is painted by connectivity using additional in-house matlab script (available after request). Volumes of the thalamic ROIs as well as FA for each individual connection are our measures of interest.
- Results presentation and paper submission. Data analysis and paper in preparation.
- Tractography Comparison Project
We are also working on a tractography comparison projectdataset, where we apply stochastic tractography to phantom, as well as test dataset.
- - Related References
- Shenton, M.E., Ngo, T., Rosenberger, G., Westin, C.F., Levitt, J.J., McCarley, R.W., Kubicki, M. Study of Thalamo-Cortical White Matter Fiber Tract Projections in Schizophrenia Using Diffusion Stochastic Tractography. Poster presented at the 46th Meeting of the American College of Neuropsychopharmacology, Boca Raton, FL, December 2007.
- Terry DP, Rausch AC, Alvarado JL, Melonakos ED, Markant D, Westin CF, Kikinis R, de Siebenthal J, Shenton ME, Kubicki M. White Matter Properties of Emotion Related Connections in Schizophrenia. Poster presented at the 2009 Mysell Poster Day, Dept. of Psychiatry, Harvard Medical School, April 2009]
- Jorge_poster.pdf Alvarado JL, Terry DP, Markant D, Ngo T, Kikinis R, Westin CF, McCarley RW, Shenton ME, Kubicki M. Study of Language-Related White Matter Tract Connections in Schizophrenia using Diffusion Stochastic Tractography. Poster presented at the 2009 Mysell Poster Day, Dept. of Psychiatry, Harvard Medical School, April 2009]
- Melonakos ED, Shenton ME, Markant D, Alvarado J, Westin CF, Kubicki M. White Matter Properties of Orbitofrontal Connections in Schizophrenia. Poster being presented at the 64th Meeting of the Society of Biological Psychiatry. Vancouver, BC. May 2009.
- Kubicki, M. Khan, U., Bobrow, L., O'Donnell, L. Pieper, S. Westin, CF., Shenton, ME. New Methods for Assessing Whole Brain DTI Abnormalities in Schizophrenia. Presentation given at the International Congress of World Psychiatric Association. Florence, Italy. April 2009.
- Kubicki, M., Markant, D., Ngo, T., Westin, CF., McCarley, RW., Shenton, ME. Study of Language Related White Matter Fiber Tract Projections in Schizophrenia Using Diffusion Stochastic Tractography. Presentation given at the International Congress of World Psychiatric Association. Florence, Italy. April 2009.
- T. Otani, M Kubicki, S. Bouix, P Nestor, A Rausch, T Asami, D, Terry, E Melonakos, K Hawley, P Pelavin, J Alvarado, A LaVenture, J Siebenthal, R McCarley, and M Shenton. White matter connections between orbitofrontal cortex and anterior cingulate cortex in shchizophrenia, Annual Meeting, Society of Biological Psychiatry, 2010.
Staffing Plan
- Sylvain and Ryan are the DBP resources charged with adapting the tools in the NA-MIC Kit to the DBP needs
- Andrew Rauch is our NAMIC RA.
- Ryan is our NAMIC software engineer. He is responsible for improving the STM (Stochastic Tractography Module), and making sure software works with STM compliant datasets.
- Polina is the algorithm core contact
- Brad is the engineering core contact
Schedule
- 10/2007 - Optimization of Stochastic Tractography algorythm for 1.5T data.
- 10/2007 - Algorythm testing on Santa Fe data set and diffusion phantom.
- 06/2008 - Optimization of Stochastic Tractography algorythm for 3T data.
- 11/2008 - Slicer 3 module prototype using python.
- 12/2008 - Slicer 3 module official release
- 12/2008 - Documentation and packaging for dissemination.
- 12/2008 - First clinical application of stochastic tractography module.
- 01/2009 - First draft of the clinical paper.
- 05/2009 - Distortion correction and nonlinear registration added to the module
- 05/2009 - Symposium on tractography, including stochastic methods at World Biological Psychiatry Symposium in Florence, Italy.
- 05/2009 - Presentation of Arcuate Fasciculus findings at World Biological Psychiatry Symposium in Florence, Italy.
- 07/2009 - Summer Programming week- work on optimizing and speeding up data processing, releasing second generation of software that includes preprocessing pipeline.
- 07/2009 - Continue working on clinical collaborative studies using stochastic tractography module.
- 12/2009 - Submission of abstracts reporting findings of several clinical studies involving stochastic tractography, including anatomical connectivity abnormalities in patients with VCFS and anatomical and functional connectivity abnormalities in schizophrenia.
- 01/2010 - AHM progress presentation.
- 05/2010 - Presentation of clinical findings at Annual Biological Psychiatry Symposium, San Francisco.
- 07/2010 - Summer Programming week- presentation of new software manual, software included in new slicer 3 release.
Team and Institute
- PI: Marek Kubicki (kubicki at bwh.harvard.edu)
- DBP2 Investigators: Sylvain Bouix, Yogesh Rathi, Ryan Ecbo
- NA-MIC Engineering Contact: Brad Davis, Kitware
- NA-MIC Algorithms Contact: Polina Gollard, MIT
Publications
In print