Difference between revisions of "Projects:RegistrationLibrary:RegLib C27"
From NAMIC Wiki
| (8 intermediate revisions by the same user not shown) | |||
| Line 3: | Line 3: | ||
[[Projects:RegistrationDocumentation:UseCaseInventory|Back to Registration Use-case Inventory]] <br> | [[Projects:RegistrationDocumentation:UseCaseInventory|Back to Registration Use-case Inventory]] <br> | ||
| − | ==<small> | + | ==<small>updated for '''v4.1'''</small> [[Image:Slicer4_RegLibLogo.png|150px]] <br>Slicer Registration Library Exampe #27: Diffusion Weighted Image Volume: align with structural reference MRI == |
=== Input === | === Input === | ||
| Line 20: | Line 20: | ||
|} | |} | ||
| − | |||
| − | |||
===Objective / Background === | ===Objective / Background === | ||
| Line 33: | Line 31: | ||
*moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 1.96 x 1.96 x 3 mm voxel size, oblique, 128 x 128 x 40 | *moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 1.96 x 1.96 x 3 mm voxel size, oblique, 128 x 128 x 40 | ||
*moving: Tensor data of DTI volume, oblique, same orientation as Baseline image. The result Xform will be applied to this volume. The original DWI has 64 directions, the extracted DTI volume has 9 scalars, i.e. 128 x 128 x 40 x 9 | *moving: Tensor data of DTI volume, oblique, same orientation as Baseline image. The result Xform will be applied to this volume. The original DWI has 64 directions, the extracted DTI volume has 9 scalars, i.e. 128 x 128 x 40 x 9 | ||
| + | |||
| + | === Overall Strategy === | ||
| + | :#Register FLAIR to T1 (affine) & resample | ||
| + | :#Resample DWI to isotropic voxel size: use T1 as size reference | ||
| + | :#Convert DWI -> DTI, incl. mask & baseline extraction | ||
| + | :#Register DTI_baseline to FLAIR (affine+nonrigid) w/o masking | ||
| + | :#Resample DTI with above transform | ||
| + | |||
| + | === Procedures === | ||
| + | *'''Phase I: register FLAIR to T1''' | ||
| + | #open the [http://wiki.slicer.org/slicerWiki/index.php/Documentation/4.1/Modules/BRAINSFit General Registration (BRAINS) module] | ||
| + | ##''Fixed Image Volume'': T1 | ||
| + | ##''Moving Image Volume'': FLAIR | ||
| + | ##Output Settings: | ||
| + | ###''Slicer BSpline Transform": none | ||
| + | ###''Slicer Linear Transform'': create new transform, rename to "Xf1_FLAIR-T1" | ||
| + | ###''Output Image Volume'': create new volume, rename to "FLAIR_Xf1" | ||
| + | ##''Regstration Phases'': check boxes for ''Rigid'' and ''Affine'' | ||
| + | ##''Main Parameters'': | ||
| + | ###''Number Of Samples'': 200,000 | ||
| + | ##Leave all other settings at default | ||
| + | ##click: Apply; runtime < 10 sec (MacPro QuadCore 2.4GHz) | ||
| + | *'''Phase II: Resample DWI to isotropic voxel size''' | ||
| + | #go to [http://wiki.slicer.org/slicerWiki/index.php/Documentation/4.1/Modules/ResampleScalarVectorDWIVolume ''ResampleScalarVectorDWIVolume'' module] (''All Modules'' menu) | ||
| + | ##''Input Volume'': "DWI", | ||
| + | ##''Output Volume'': "create new Diffusion Weighted Volume", rename to "DWI_iso" | ||
| + | ##''Reference Volume'': T1 | ||
| + | #Click: ''Apply'' | ||
| + | #save new DWI_iso to disk | ||
| + | #depending on RAM of your machine, consider deleting the original DWI node | ||
| + | *'''Phase III: DWI -> DTI''' | ||
| + | #open [http://wiki.slicer.org/slicerWiki/index.php/Documentation/4.1/Modules/DiffusionTensorEstimation "Diffusion Tensor Estimation" module] (menu: Diffusion:DiffusionWeightedImages: DiffusionTensorEstimation) | ||
| + | ##''Input DWI Volume'': DWI_iso, | ||
| + | ##''Output DTI Volume'': create new, rename to "DTI_iso" | ||
| + | ##''Output Baseline Volume'': create new, rename to "DWI_iso_baseline" | ||
| + | #Click: Apply | ||
| + | *'''Phase IV: Register DTI (unmasked)''' | ||
| + | #open the [http://wiki.slicer.org/slicerWiki/index.php/Documentation/4.1/Modules/BRAINSFit General Registration (BRAINS) module] | ||
| + | ##''Fixed Image Volume'': FLAIR_Xf1 | ||
| + | ##''Moving Image Volume'': DWI_iso_baseline | ||
| + | ##Output Settings: | ||
| + | ###''Slicer BSpline Transform": create new transform, rename to "Xf2_DTI-T1" | ||
| + | ###''Slicer Linear Transform'': none | ||
| + | ###''Output Image Volume'': create new volume, rename to DWI_iso_baseline_Xf2 | ||
| + | ##''Regstration Phases'': check boxes for ''Rigid'', ''Affine'' and ''BSpline'' | ||
| + | ##''Main Parameters'': | ||
| + | ###''Number Of Samples'': 200,000 | ||
| + | ###''B-Spline Grid Size'': 5,5,3 | ||
| + | ##Leave all other settings at default | ||
| + | ##click: Apply; runtime <1 min (MacPro QuadCore 2.4GHz) | ||
| + | *'''Phase V: Resample DTI''' | ||
| + | #Open the [http://wiki.slicer.org/slicerWiki/index.php/Documentation/4.1/Modules/ResampleDTI ''Resample DTI Volume'' module] (under ''All Modules'' menu; note this is distinct from the ResampleScalarVectorDWIVolume used above) | ||
| + | ##''Input Volume'': DTI_iso | ||
| + | ##''Output Volume'': create new DTI Volume, rename to ''DTI_iso_Xf2'' | ||
| + | ##''Reference Volume'': T1 | ||
| + | ##''Transform Node'': select "Xf2_DTI-T1'' created in phase 4 above | ||
| + | ##check box: ''displacement'' | ||
| + | #leave all other settings at defaults | ||
| + | #Click Apply; runtime ~ 3 min. | ||
| + | #set T1 or FLAIR as background and the new ''DTI_Xf2'' volume as foreground | ||
| + | #Move fade slider to see DTI overlay onto the structural image | ||
=== Registration Results=== | === Registration Results=== | ||
| Line 42: | Line 101: | ||
*Data | *Data | ||
**[[Media:RegLib_C27_Data.zip |'''RegLib_C27_Data''' DWI, DTI, T1, FLAIR, Presets & solution transforms <small> (NRRD files, zip file 145 MB) </small>]]''' | **[[Media:RegLib_C27_Data.zip |'''RegLib_C27_Data''' DWI, DTI, T1, FLAIR, Presets & solution transforms <small> (NRRD files, zip file 145 MB) </small>]]''' | ||
| − | * | + | *Documentation (for Slicer Version 3.6, but procedure is the same) |
| − | |||
| − | |||
| − | |||
| − | |||
**[[Media:RegLib_C27_Tutorial.ppt|'''RegLib_C27_Tutorial.ppt''' step-by-step tutorial Powerpoint<small> (.ppt file 2.6MB) </small>]]''' | **[[Media:RegLib_C27_Tutorial.ppt|'''RegLib_C27_Tutorial.ppt''' step-by-step tutorial Powerpoint<small> (.ppt file 2.6MB) </small>]]''' | ||
**[[Media:RegLib_C27_Tutorial.pdf|'''RegLib_C27_Tutorial.pdf''' step-by-step tutorial PDF<small> (.pdf file 3.8MB) </small>]]''' | **[[Media:RegLib_C27_Tutorial.pdf|'''RegLib_C27_Tutorial.pdf''' step-by-step tutorial PDF<small> (.pdf file 3.8MB) </small>]]''' | ||
Latest revision as of 20:06, 4 May 2012
Home < Projects:RegistrationLibrary:RegLib C27Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
updated for v4.1 
Slicer Registration Library Exampe #27: Diffusion Weighted Image Volume: align with structural reference MRI
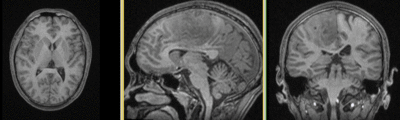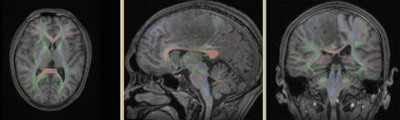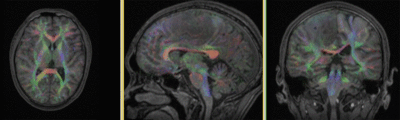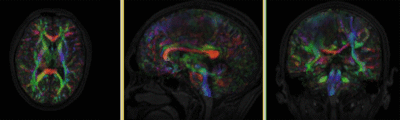
Input
|
|
|
|
|
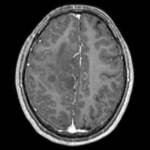
| fixed image/target T1 |
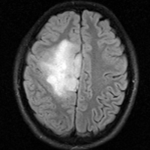
fixed image/target FLAIR |
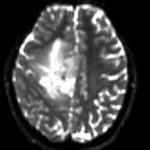
moving image 2a DTI baseline |
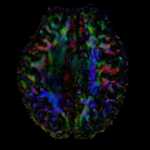
moving image 2b DTI tensor |
Objective / Background
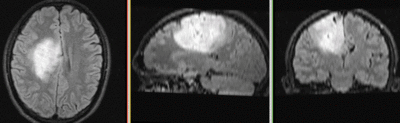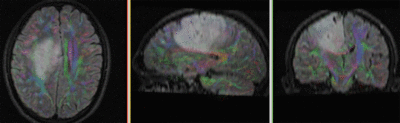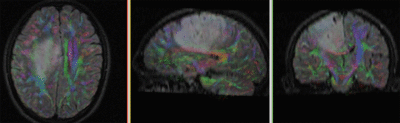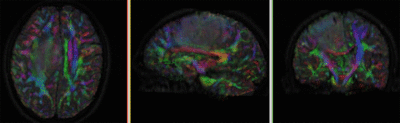
This is a typical example of DTI processing. Goal is to align the DTI image with a structural scan that provides accuracte anatomical reference. The DTI contains acquisition-related distortion and insufficient contrast to discern anatomical detail. For treatment planning and evaluation, location of functionally critical fiber tracts relative to the pathology is sought.
Keywords
MRI, brain, head, intra-subject, DTI, DWI
Input Data
- reference/fixed : T1 axial, 0.98 x 0.98 x 1 , 192 x 256 x 176
- reference/fixed : FLAIR axial, 0.4mm resolution in plane, 4mm slices, 448 x 512 x 30
- moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 1.96 x 1.96 x 3 mm voxel size, oblique, 128 x 128 x 40
- moving: Tensor data of DTI volume, oblique, same orientation as Baseline image. The result Xform will be applied to this volume. The original DWI has 64 directions, the extracted DTI volume has 9 scalars, i.e. 128 x 128 x 40 x 9
Overall Strategy
- Register FLAIR to T1 (affine) & resample
- Resample DWI to isotropic voxel size: use T1 as size reference
- Convert DWI -> DTI, incl. mask & baseline extraction
- Register DTI_baseline to FLAIR (affine+nonrigid) w/o masking
- Resample DTI with above transform
Procedures
- Phase I: register FLAIR to T1
- open the General Registration (BRAINS) module
- Fixed Image Volume: T1
- Moving Image Volume: FLAIR
- Output Settings:
- Slicer BSpline Transform": none
- Slicer Linear Transform: create new transform, rename to "Xf1_FLAIR-T1"
- Output Image Volume: create new volume, rename to "FLAIR_Xf1"
- Regstration Phases: check boxes for Rigid and Affine
- Main Parameters:
- Number Of Samples: 200,000
- Leave all other settings at default
- click: Apply; runtime < 10 sec (MacPro QuadCore 2.4GHz)
- Phase II: Resample DWI to isotropic voxel size
- go to ResampleScalarVectorDWIVolume module (All Modules menu)
- Input Volume: "DWI",
- Output Volume: "create new Diffusion Weighted Volume", rename to "DWI_iso"
- Reference Volume: T1
- Click: Apply
- save new DWI_iso to disk
- depending on RAM of your machine, consider deleting the original DWI node
- Phase III: DWI -> DTI
- open "Diffusion Tensor Estimation" module (menu: Diffusion:DiffusionWeightedImages: DiffusionTensorEstimation)
- Input DWI Volume: DWI_iso,
- Output DTI Volume: create new, rename to "DTI_iso"
- Output Baseline Volume: create new, rename to "DWI_iso_baseline"
- Click: Apply
- Phase IV: Register DTI (unmasked)
- open the General Registration (BRAINS) module
- Fixed Image Volume: FLAIR_Xf1
- Moving Image Volume: DWI_iso_baseline
- Output Settings:
- Slicer BSpline Transform": create new transform, rename to "Xf2_DTI-T1"
- Slicer Linear Transform: none
- Output Image Volume: create new volume, rename to DWI_iso_baseline_Xf2
- Regstration Phases: check boxes for Rigid, Affine and BSpline
- Main Parameters:
- Number Of Samples: 200,000
- B-Spline Grid Size: 5,5,3
- Leave all other settings at default
- click: Apply; runtime <1 min (MacPro QuadCore 2.4GHz)
- Phase V: Resample DTI
- Open the Resample DTI Volume module (under All Modules menu; note this is distinct from the ResampleScalarVectorDWIVolume used above)
- Input Volume: DTI_iso
- Output Volume: create new DTI Volume, rename to DTI_iso_Xf2
- Reference Volume: T1
- Transform Node: select "Xf2_DTI-T1 created in phase 4 above
- check box: displacement
- leave all other settings at defaults
- Click Apply; runtime ~ 3 min.
- set T1 or FLAIR as background and the new DTI_Xf2 volume as foreground
- Move fade slider to see DTI overlay onto the structural image
Registration Results
Download
- Data
- Documentation (for Slicer Version 3.6, but procedure is the same)
Discussion: Registration Challenges
- The DTI contains acquisition-related distortions (commonly EPI acquisitions) that can make automated registration difficult.
- the two images often have strong differences in voxel sizes and voxel anisotropy. If the orientation of the highest resolution is not the same in both images, finding a good match can be difficult.
- there is widespread and extensive pathology in the right hemisphere that might affect the registration if contrast is different between the baseline and structural reference scan.
Discussion: Key Strategies
- because the pathology appears similar in the FLAIR as in the DTI baseline, we choose the FLAIR as reference
- masking is likely necessary to obtain good results.
- in this example the initial alignment of the two scans is pretty good already. No initial affine alignment is needed.
- these two images are not too far apart initially, so we reduce the default of expected translational misalignment
- because speed is not that critical, we increase the sampling rate from the default 2% to 15%.