2016 Summer Project Week/Brain atlas combining histology and MRI
From NAMIC Wiki
Home < 2016 Summer Project Week < Brain atlas combining histology and MRI
Key Investigators
- Fernando Pérez García (Institute of the Brain and Spine, France)
- Sara Fernandez Vidal (Institute of the Brain and Spine, France)
- Sonia Pujol (BWH)
- Steve Pieper
- Csaba Pinter
- Andras Lasso
Project Description
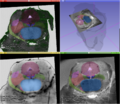
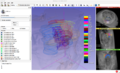
Multimodal 3D visualization is important during the construction of a 3D histology / MRI atlas. Also, the interface must be very intuitive so that the user can easily trace the many contours needed for a precise 3D reconstruction.
We have chosen 3D Slicer as our development platform because of its flexibility and its huge potential for multimodal 3D visualization. Our goal is to develop an integrated, intuitive extension for manual tracing of contours on stained histological slices that have been coregistered to a high resolution MRI.
| Objective | Approach and Plan | Progress and Next Steps |
|---|---|---|
|
|
|