NAMIC Wiki:DTI:Nrrd format
Contents
- 1 Minimal math background on DWI
- 2 NRRD format usage for NAMIC DWI volumes
- 3 Learning by real-world example: a Dartmouth DWI header
- 4 Understanding necessary NRRD fields
- 5 Key/Value pair convention for DWI
- 6 Describing DWIs with different b-values
- 7 Pseudo-code algorithm for parsing DWI key/value pairs
- 8 Software support for NRRD DWIs
- 9 Synthetic helical DWI volume for coordinate stress-testing
Minimal math background on DWI
In the standard diffusion-weighted image (DWI) acquisition for tensor estimation, the assumption is that the measured image intensity [math]A_i[/math] is modeled by:
[math]A_i = A_0 \exp(-b\, \mathbf{g}_i^T \mathbf{D} \mathbf{g}_i)[/math]
[math]= A_0 \exp(-b\, ( \mathbf{g}_{x_i} \mathbf{g}_{x_i} \mathbf{D}_{xx} + 2 \mathbf{g}_{x_i} \mathbf{g}_{y_i} \mathbf{D}_{xy} + 2 \mathbf{g}_{x_i} \mathbf{g}_{z_i} \mathbf{D}_{xz} + \mathbf{g}_{y_i} \mathbf{g}_{y_i} \mathbf{D}_{yy} + 2 \mathbf{g}_{y_i} \mathbf{g}_{z_i} \mathbf{D}_{yz} + \mathbf{g}_{z_i} \mathbf{g}_{z_i} \mathbf{D}_{zz} ))[/math]
where [math]A_0[/math] is the non-diffusion-weighted T2 image intensity, [math]b[/math] is the diffusion weighting parameter, [math]\mathbf{g}[/math] is the diffusion-sensitizing gradient direction, and [math]\mathbf{D}[/math] is the diffusion tensor. The measured image gets dimmer according to how much diffusion happened along the gradient direction. Tensors can be estimated from one or more measurements of [math]A_0[/math], and a set of [math]A_i[/math] for a set of at least 6 gradient directions [math]\mathbf{g}_i[/math].
In some experiments more accuracy may be obtained in the tensor estimate by taking into account the diffusion weighting induced by the imaging gradients. This information is less commonly known, but may be calculated from the specifics of the pulse sequence. In this case, the contribution of the coefficients of [math]\mathbf{D}[/math] to the measured DWI is represented as
[math]A_i\,[/math] [math]= A_0 \exp(-b\,\mathbf{B} \mathbf{:} \mathbf{D})[/math] [math]= A_0 \exp(-b\,\mathrm{trace}(\mathbf{B}^T \mathbf{D}))[/math] [math]= A_0 \exp(-b\, ( \mathbf{B}_{{xx}_i} \mathbf{D}_{xx} + 2 \mathbf{B}_{{xy}_i} \mathbf{D}_{xy} + 2 \mathbf{B}_{{xz}_i} \mathbf{D}_{xz} + \mathbf{B}_{{yy}_i} \mathbf{D}_{yy} + 2 \mathbf{B}_{{yz}_i} \mathbf{D}_{yz} + \mathbf{B}_{{zz}_i} \mathbf{D}_{zz} ))[/math]
where [math]\mathbf{B}[/math] is known from the acquisition. In the simple case [math]\mathbf{B} = \mathbf{g}\mathbf{g}^T[/math].
The main purpose of a file format for DWI is to record all the information necessary to unambiguously reconstruct tensors from the measured images. This includes:
- b-value
- gradient directions (or full B-matrices, when known)
- the "measurement frame": relationship between the coordinate frame in which the gradient coefficients are expressed, and the physical coordinate frame in which image orientation is defined
- order in which the different DWI values were acquired (e.g. slice interleaved vs. volume interleaved)
- anatomical location of all the images (as with any acquisition)
NRRD format usage for NAMIC DWI volumes
The NRRD file format, combined with a particular convention for using key/value pairs in the format, can be used to completely represent the necessary information about a DWI image volume, its anatomical orientation, and all the DWI-specific acquisition parameters that determine how the image values are used to estimate diffusion tensors.
This page about DICOM and DWI can serve as a place where information about DICOM support for DWI and DTI data can be collected and updated.
The application of the NRRD format for representing DWI datasets, is motivated by the following situation:
- Even though there is nominally a standard for how scanners should store the DWI parameters, current scanners usually don't comply with the standard, and instead have disparate and manufacturer-specific ways of using DICOM tags
- Even when gradient directions can be learned from DICOM tags, there is no record of the measurement frame, which is necessary for fully-informed processing of the data (it should be noted that in the "standard", the measurement frame is the identity)
- Even when DWI-related tags are defined and used, there have been instances where the information is not set correctly (for example in research protocols which are modifications of manufacturer-supplied sequences)
- Many older datasets, still necessary for research, are not in DICOM format, or in any widely used format
This page documents the current "standard" for using NRRD for DWI datasets in NAMIC. It is intended as a clarification/restatement of the older information on this page, with a few limited enhancements:
- In the 4 axes of the image, the non-spatial axis (for DWI values) is distinguished from the 3 spatial axes by either the "list" [kind], or the "vector" kind.
- The non-spatial axis can appear anywhere in the axis ordering, not just as the slowest axis.
- The way that images with different b-values are represented has been fixed/clarified: If the b-value is scaled by a factor of X, the length of the corresponding gradient vector should be scaled by a factor of sqrt(X) (not X, as a previous example used).
Learning by real-world example: a Dartmouth DWI header
The simplest way to show how the NRRD format fields (see below) are used to represent the DWI information is by example. Specifically, the NRRD DWI headers demonstrate everything involved. Here is a breakdown of the header for NAMIC01dti-anon, with some re-ordering of the lines for the sake of explanation.
NRRD0005 # Complete NRRD file format specification at: # http://teem.sourceforge.net/nrrd/format.html content: NAMIC01
This is the format "magic" given the version of the NRRD format used, a comment for how to find the format info, and an identifier of which dataset this is. The all-important "measurement frame" field did not exist in NRRD0004, and is new to NRRD0005.
dimension: 4 sizes: 256 256 36 14 centerings: cell cell cell none thicknesses: NaN NaN 3 NaN kinds: space space space list
These fields define the logical structure of the image data as a 4D array, as well as some information about pixel centering (node-centered versus cell-centered), pixel thickness (a fairly medical-image-specific concept).
The "kinds" field identifies which of the four axes is for storing the DWI image values, as opposed to the 3 spatial image domain axes. The single non-spatial axis is identified by either "list" or "vector" (the latter being allowed to be compatible with how ITK writes out non-scalar NRRDs by default).
The location of the non-spatial axis among the spatial axes determines the interleaving of the pixel data (which values you see when as you traverse through all the image values in memory address order) For example:
- "kinds: space space space list": volume interleaved: first you go through all the slices for one gradient direction, and then you go through the slices for a second gradient direction.
- "kinds: space space list space": slice interleaved: for one 2-D slice location you go through the DWI values, before you move on to the next physical slice.
- "kinds: list space space space": pixel interleaved: you go through the DWI values for each pixel before moving on to the values for the next pixel.
Previous versions of the NAMIC NRRD DWI format suggested that the "list" axis always be on the slowest axis (volume interleaved). This is an undesirable restriction, because it prevents the NRRD header from referring to existing data files for which the data is something other than volume interleaved.
type: short endian: little encoding: raw data file: S4.%03d 1 504 1 2 byte skip: -1
This is all the information about how the raw image values are stored in the multiple image files. NOTE that this only works because in this case there is an alphanumeric ordering of the files that is consistent with the raster ordering of data. If there is interleaving, for example, you can use the "data file: LIST" field and end the header with a complete list of all the constituent files. As these are 2-D image slices composing a 4-D volume, the datafile dimension ("2") must be specified as the last parameter to the [data file] field.
space: right-anterior-superior space directions: (-0.9375,0,0) (0,-0.9375,0) (0,0,-3) none space units: "mm" "mm" "mm" space origin: (125.00,124.10,79.3)
These fields define image orientation. This example uses RAS coordinates, but it is also possible to use DICOM's LPS. The "space directions" field defines, one column vector at a time, the transform from IJK raster image coordinates (I==fastest;K==slowest) to RAS anatomical image coordinates:
[math]\begin{bmatrix} \mathrm{R} \\ \mathrm{A} \\ \mathrm{S} \end{bmatrix} = \begin{bmatrix} -0.9375 & 0.0 & 0.0 & 125.0 \\ 0.0 & -0.9375 & 0.0 & 124.1 \\ 0.0 & 0.0 & -3.0 & 79.3 \end{bmatrix} \begin{bmatrix} \mathrm{I} \\ \mathrm{J} \\ \mathrm{K} \\ 1.0 \end{bmatrix}[/math]
measurement frame: (0,-1,0) (1,0,0) (0,0,-1)
This defines, one column vector at a time, the matrix which transforms the (X,Y,Z) coordinates used for listing the diffusion-sensitizing gradient directions (below), into whatever space is named by "space", in this case RAS anatomical image coordinates:
[math]\begin{bmatrix} \mathrm{R} \\ \mathrm{A} \\ \mathrm{S} \end{bmatrix} = \begin{bmatrix} 0 & 1 & 0 \\ -1 & 0 & 0 \\ 0 & 0 & -1 \end{bmatrix} \begin{bmatrix} \mathrm{X} \\ \mathrm{Y} \\ \mathrm{Z} \end{bmatrix}[/math]
modality:=DWMRI DWMRI_b-value:=800 DWMRI_gradient_0000:= 0 0 0 DWMRI_NEX_0000:=2 DWMRI_gradient_0002:= -0.8238094 -0.4178235 -0.3830949 DWMRI_gradient_0003:= -0.5681645 0.5019867 -0.6520725 DWMRI_gradient_0004:= 0.4296590 0.1437401 0.8914774 DWMRI_gradient_0005:= -0.0482123 0.6979894 0.7144833 DWMRI_gradient_0006:= 0.8286872 -0.0896669 -0.5524829 DWMRI_gradient_0007:= 0.9642489 -0.2240180 0.1415627 DWMRI_gradient_0008:= -0.1944068 0.9526976 -0.2336092 DWMRI_gradient_0009:= 0.1662157 0.6172332 -0.7690224 DWMRI_gradient_0010:= -0.3535898 -0.9178798 -0.1801968 DWMRI_gradient_0011:= -0.7404186 -0.5774342 0.3440203 DWMRI_gradient_0012:= -0.2763061 0.0476582 0.9598873 DWMRI_gradient_0013:= 0.6168819 -0.7348858 -0.2817793
These define the scalar diffusion weighting b-value, and the diffusion-weighting information for all the image values acquired per-pixel: first the two baseline (non-diffusion-weighted) images, and then the 12 DWIs. NOTE: there is no requirement that the baseline image be first. It could be last, or appear in multiple places, or appear not at all.
Understanding necessary NRRD fields
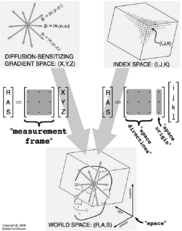
The diagram at right illustrates the "space", "space origin", "space directions", and "measurement frame" fields of the NRRD header, which serve the important role of defining the relationship between the IJK image coordinates, the XYZ diffusion-sensitizing gradient coordinates, and the RAS anatomical coordinates.
An introduction to how NRRD represents raster data [is available here]. This page is useful because it explains how what is physically a 3D image is stored as a 4D array whenever there are multiple values per pixel. The idea of "fast" and "slow" axes is important to understand, as well as the terminology of (image) "dimension" and (axis) "size".
The full definition of the NRRD file format [is available here]. This document is somewhat lengthy, but it makes every effort to be unambiguous and helpful in explaining what the various NRRD fields are. The fields required for the NAMIC DWI NRRD format convention include:
- Image resolution and basic geometrical properties handled by [dimension], [sizes], [thicknesses], and [centerings] fields. Note:
- The "dimension" field has to precede all the pre-axis fields
- DWI volumes will use "dimension: 4", and the "sizes:" field has to give the correct logical axis sizes of the 4D image array. The total number of baseline and diffusion-weighted images will determine one of the four axis sizes.
- Numerical data representation and file info handled by [type], [encoding], [endian], and [data file] fields. The "data file:" field allows the data to be in single or multiple separate files.
- Basic meta-information about value attributes is handled by [content] and [kinds] fields. The "kinds" field is important because it is how you mark which of the four array axes is for the DWI values, and which axes are the spatial domain of the image. Specifically, the "list" or "vector" kind identifies the axis for DWI values.
- Anatomical image orientation is handled by [space], [space origin], [space directions], and [space units] fields. This is the information that in DICOM is handled with "Image Position (Patient)" and "Image Orientation (Patient)".
- An important capability of the NRRD format is its explicit representation of the [measurement frame]. This clarifies what continues to be a confusing aspect of working with DWI data: the relationship between the coordinate frame in which the coefficients of the diffusion gradients (or the diffusion tensor) are measured, and the coordinate frame of the image itself. The "measurement frame" field identifies the mapping from coordinates associated with the diffusion gradients/tensors to the world space coordinates associated with the image orientation.
Key/Value pair convention for DWI
This is how the DWI-specific information is stored in the NRRD header, which by itself doesn't know anything about DWI.
- "modality:=DWMRI" : This key/value pair explicitly indicates that the image is a diffusion-weighted MRI scan, and it implies that all of the following key/value pairs are also set in the header:
- "DWMRI_b-value:=b " : This key/value pair gives the (scalar) diffusion-weighting value, in units of s/mm^2. Example: "DWMRI_b-value:=1000". As described below, the effective magnitude of diffusion-weighting for each DWI is determined with some simple calculations based on the individual per-DWI gradient directions or B-matrices.
- For every index position NNNN along the DWI axis (whichever is the non-spatial axis identified by the "list" or "vector" kind field), either "DWMRI_gradient_NNNN:=x y z " or "DWMRI_B-matrix_NNNN:=xx xy xz yy yz zz " must be given (except if "DWMRI_NEX_NNNN:= M " is used).
- "DWMRI_NEX_NNNN:=M " means that the information provided for image NNNN (either gradient or B-matrix) is the same as for image NNNN+1, NNNN+2, up to and including NNNN+M-1. So there are M repeated acquisitions of the image starting at index NNNN, and the next M-1 gradient (or B-matrix) keys are elided.
- For simplicity, it is probably best to have all DWMRI_gradient or all DWMRI_B-matrix keys, rather than some intermixing of both
- NNNN should start at 0000, and go up to numImg-1, always using 4 digits to represent the index (as if by the "%04d" format string to sprintf()).
- The gradient "direction" corresponding to a truly non-diffusion-weighted reference image is "0 0 0", and the B-matrix is "0 0 0 0 0 0". Example: "DWMRI_gradient_0000:=0 0 0".
- When specified by "DWMRI_B-matrix_NNNN", the B-matrix off-diagonal entries are NOT pre-multiplied by 2. So, the actual DWI value for B-matrix "Bxx Bxy Bxz Byy Byz Bzz" and diffusion tensor D is modeled by the second equation above.
In addition to all the info above, there is an implicit normalization of the gradients or B-matrices. This permits non-unit length gradients to be stored in a manner which corresponds (from a human-readable standpoint) to their conventional expression, such as the gradient set used in the Harvard/Brockton VA data: {(1,1,0), (0,1,1), (1,0,1), (0,1,-1), (1,-1,0), (-1,0,1)}).
The implicit normalization is implementing by dividing all the gradients (or B-matrices) by the maximal gradient (or B-matrix) magnitude. The magnitude of a gradient direction vector is determined by the usual L^2 norm, and the magnitude of a B-matrix is via the [Frobenius Norm]. It is after this magnitude rescaling that the nominal b-value (given via "DWMRI_b-value:=b") applies.
Describing DWIs with different b-values
As currently defined, the key/value pair convention above only names a single b-value. It is possible, however, to describe acquisitions with different b-values, by varying the magnitudes of the gradients (or B-matrices). Note from the math background above that if the (L2) length of g scaled by x, then the magnitude of the exponent is scaled by x2.
This determines how to scale the gradient lengths to convey different b-values. If the nominal b-value is 1000 (via DWMRI_b-value:=1000), then to represent a DWI with b=500, use a gradient vector (or B-matrix) who's norm is sqrt(1/2) the norm for the b=1000 DWI.
Here is a complete (and real-world) example to demonstrate this capability. The volume is 128x128x59, and for every voxel there are 13 values, 1 non-diffusion-weighted, and then two sets of DWIs using the same 6 gradients, first at b=500 s/mm2, then at b=1000 s/mm2.
NRRD0005 # Complete NRRD file format specification at: # http://teem.sourceforge.net/nrrd/format.html content: 0002 type: short dimension: 4 sizes: 128 128 59 13 centerings: cell cell cell none kinds: space space space list space: left-posterior-superior thicknesses: NaN NaN 2.2 NaN space origin: (-128,-142.23729,99.732201) space directions: (2,0,0) (0,2,0) (0,0,-2.199997) none space units: "mm" "mm" "mm" endian: little encoding: raw data file: SerieMR-0002-%04d.dcm 1 767 1 2 byte skip: -1 measurement frame: (-1,0,0) (0,1,0) (0,0,1) modality:=DWMRI DWMRI_b-value:=1000 DWMRI_gradient_0000:= 0 0 0 DWMRI_gradient_0001:= 0.707107 0.0 0.707107 DWMRI_gradient_0002:= -0.707107 0.0 0.707107 DWMRI_gradient_0003:= 0.0 0.707107 0.707107 DWMRI_gradient_0004:= 0.0 0.707107 -0.707107 DWMRI_gradient_0005:= 0.707107 0.707107 0.0 DWMRI_gradient_0006:= -0.707107 0.707107 0.0 DWMRI_gradient_0007:= 1 0 1 DWMRI_gradient_0008:= -1 0 1 DWMRI_gradient_0009:= 0 1 1 DWMRI_gradient_0010:= 0 1 -1 DWMRI_gradient_0011:= 1 1 0 DWMRI_gradient_0012:= -1 1 0
Pseudo-code algorithm for parsing DWI key/value pairs
... to be finished ...
Software support for NRRD DWIs
- Teem: Examples of using Teem tools for DWI processing: Using the unu and tend command-line tools for initial processing, and Deft for visualization.
- Slicer: Slicer (2.6 and later) support nrrd data through the nrrd Reader. The supported data types are: Scalar 3D images (grayscale and color), DWI images, Tensors and general 4D arrays. A general 4D array will be represented as a scalar 3D volumes with multiple components.
- Loading the data: Add Volume -> Properties: Nrrd Reader -> Choose the file to load.
- Converting DWI to Tensor: Go to DTMRI Module -> Conv Tab -> Choose the Volume node -> Convert. The information about gradients and reference frames are carried from the file to do the conversion.
- Visualizing Tensor data: Load tensor nrrd file -> Go to DTMRI Module -> Glyph -> Choose tensor node -> Display Glyphs On.
- ITK: can read both DWIs and DTIs from NRRD, as long as you correctly create the reader!
- ... DWIs using fixed-length vectors ...
- ... DWIs using VectorImage (learned at run-time) ...
- ITK can estimate tensors with the ...???... filter
- DTI volumes can be read via:
typedef itk::DiffusionTensor3D<float> PixelType;
typedef itk::Image<PixelType, 3> myImage
itk::ImageFileReader<myImage>::Pointer reader
= itk::ImageFileReader<myImage>::New();
reader->SetFileName("filename.nrrd");
reader->Update();
myImage::Pointer image = reader->GetOutput();
Synthetic helical DWI volume for coordinate stress-testing
The command-line tools in Teem can be used to simulated DWI images, and store them with a NRRD header. Because of the known structure of the data, the DWI dataset can be used to debug a given program's handling of the various coordinate systems (image IJK, anatomical RAS/LPS, scanner XYZ).