Projects:DTIStochasticTractography
Back to NA-MIC Collaborations, NAC Algorithms, MIT Algorithms, Harvard DBP1
DTI Stochastic Tractography
Stochastic Tractography is a Bayesian approach to estimating nerve fiber tracts from DWMRI (Diffusion Weighted Magnetic Imaging) images. The Bayesian framework provides a measure of confidence regarding the estimated tracts. This measure of confidence allows the algorithm to generate tracts which pass through regions with uncertain fiber directions, revealing more details about structural connectivity than non-Bayesian tractography algorithms.
Description
Magnetic Resonance Imaging (MRI) is a valuable imaging modality for studying the brain in-vivo. We can use use MRI to differentiate between tissue types, which is valuable for anatomical studies. However, anatomical MRI provides a homogeneous image of white matter making it difficult to characterize white matter fiber tracts which pass through this region. Diffusion Weighted Magnetic Resonance Imaging (DWMRI) provides information about the diffusion of water molecules in the brain. DWMRI images can be used to construct a DTI data set which provides a complete description of water diffusion.
Researchers have hypothesized that white matter abnormalities may underlie some neurological conditions. For instance, the neurological disease schizophrenia by is characterized by its behavioral symptoms, which include auditory hallucinations, disordered thinking and delusion[6]. Studies have suggested that these behavioral symptoms have some connection with the neuroanatomical abnormalities observed in schizophrenia patients. Using DTI, Researchers can noninvasively investigate the relationship between brain white matter abnormalities and schizophrenia by using DTI.
We can visualize DTI data sets using a number of methods. DTI provides information about the diffusion of water at each voxel, or volume element in the form of diffusion tensors. A popular technique to visualize these diffusion tensors is to draw fiber tracts which utilize the diffusion information across many voxels. This technique is known as DTI White Matter Tractography.
One possible method to perform tractography is to draw tracts which are oriented along the direction of maximal water diffusion of the voxels they pass through[1]. However, this method does not provide information about the uncertainty of the generated tracks due to noise or insufficient spatial resolution. Probabilistic white matter tractography addresses this problem by performing tractography under a probabilistic framework and provides a metric for assessing the uncertainty of generated fiber tracts. Several mathematical formulations of probabilistic tractography have existed for some time with the earliest being Behren's [3].
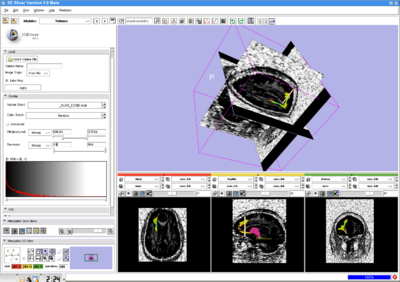
Ultimately the success of the algorithm will depend on its use in the research community. To this end, we have created a complete user interface to support the algorithm. This interface will be integrated into the popular 3D Slicer medical image visualization program. Additionally, the algorithm will be implemented within the ITK medical image analysis toolkit. ITK provides a standardized programming interface for a large collection of medical image processing algorithms which enable application developers to quickly incorporate the algorithms into new applications.
Progress
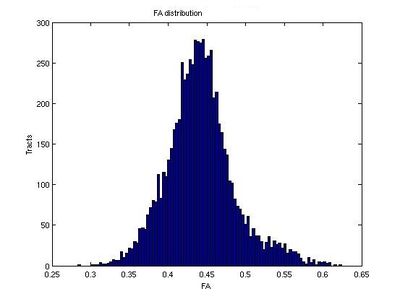
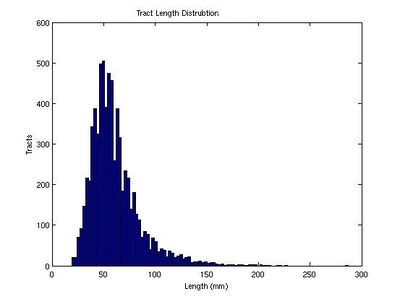
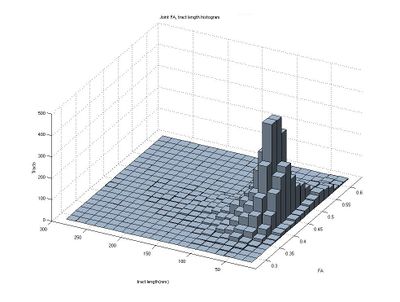
Here we estimate the distribution of Tract-Average FA and tract lengths (in mm) for tracts which originate from the right internal capsule and progress toward the prefrontal cortex.
NAMIC Software
A 3D Slicer module has been created using the command line module interface introduced in 3D Slicer version 3.
The software developed in this project includes:
- New multithreaded ITK Filter (itkStochasticTractographyFilter)
- 3D Slicer Command Line Module
- Allows the algorithm to be executed without 3D Slicer.
Key Investigators
- MIT: Tri Ngo, Polina Golland
- BWH/Harvard: NAC: Carl-Fredrik Westin
- BWH/Harvard: DBP1: Marek Kubicki
- Kitware: Brad Davis
Publications
In Print
Links
- Brigham and Women's Hospital. 3d slicer medical visualization and processing environment for research. http://www.slicer.org/.
- Insight Software Consortium. National library of medicine insight segmentation and registration toolkit(itk). http://www.itk.org/.
Project Week Results: Jan 2007