Leadership:Main
From NAMIC Wiki
Home < Leadership:Main
Back to NA-MIC Cores
Introduction
|
The National Alliance for Medical Image Computing,NA-MIC, is a network of peers consisting of computer scientists, engineers and biomedical researchers. NA-MIC has multiple goals which include:
The main role of the leadership of this enterprise is to set the long term and short term goals for the Alliance. This requires close consultation and coordination with and among the participants of NA-MIC. A secondary role of the leadership core is to present NA-MIC to the outside world, including NIH, the biomedical and medical image computing communities. |
NA-MIC Activities | |
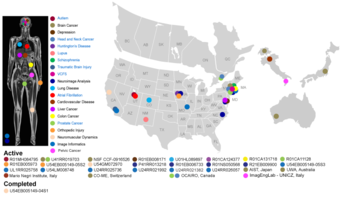
| NA-MIC is a National Alliance with funded activities in a variety of disease processes, geographic locations and, anatomic regions. | ||
Regular Meetings
- Weekly Engineering Tcons
- Monthly Algorithm Tcons
- Regular phone calls with core PI's
Activities
2014
- Juli 1 2014: 2014 Post NCBC Funding
- May 1: Queens University School of Computing Distinguished Seminar. See here for the presentation.
- March 5-9: Keynote speech at GSDM Inauguration Symposium, Tokyo, Japan.
2013
- December 16-17: Invited talk at the Medical University of South Carolina, Charleston, SC.
- December 1: RSNA 2013
- November 28, 2013: The CURAC society awards the inaugural honorary membership to Ron Kikinis during the 12th annual meeting of that society in Innsbruck, Austria. (see also 1, 2). See here for the presentation.
- October: after the 2013 DTI Challenge
- September 30: 3D Slicer invited seminar, Changchun, China.
- September 29: 3D Slicer invited workshop, PLA Hospital, Beijing, China.
- September 25: Special Session (Panel Discussion):"How do our experts see the future of medical imaging and computer assisted interventions?", Nagoya, JP. Rons slides here.
- September 22: MICCAI 2013 DTI Tractography Challenge on Peritumoral White Matter Anatomy for Neurosurgical Decision-Making
- September 5-7: Invited Keynote Lecture at the international SMIT 2013 conference. Baden-Baden, Germany.
- September 2: Invited Plenary Lecture at the opening event of the GMDS conference. Lübeck, Germany.
- July 29-30: Workshop on Enhancing Training for Biomedical Big Data, Rockville, MD, (Program)
- June 17-21: Summer Project Week at MIT
- April 10-11: Slicer Ultrasound Workshop, Iwate, Japan.
- April 9: Slicer Workshop, Tokyo, Japan.
- January 7-11: AHM, EAB, Winter Project Week in Salt Lake City
2012
- November 8-9 2012: NCBC showcase event in Washington DC: PDF of the overview slides

|
|
- September: Raise the visibility of MIC: Wikipedia page, call for identifying MIC grants submitted to NIH
- August 31: Tcon for planning Nov NIH event
- May: Visit to Australia with presentations in Sydney, Melbourne, Perth, Australia.

|

|
2011
- November 27: Slicer 4 release at RSNA, Chicago, IL.
- November 16: Opening lecture at CASEIB, Carceres, Spain.
- November 6: Slicer course at AAGL, Ft. Lauderdale, FL.
- October 20: Slicer 4 Documentation Sprint
- October 14-15: Algorithm Core retreat
- October 12-13: IGT workshop,Washington DC.
- October 7: Presentation at UBC
- September 18: DTI challenge at Miccai
- August 24: Dicom ITK discussion
- June 15: Slicer Workshop, Biomedical Imaging Research Centre at the University of Western Ontario
- May 11-12: iDASH Symposium: Integrating Data for Analysis, Anonymization, and Sharing Namic presentation on day 1). The Great Hall, University of California, San Diego, CA.
- April 23-30: Roadtrip in Japan.
- April 22:Working meeting with the AFIB DBP, Salt lake City, UT.
- April 5: Presentation, JHU. See here for the entire event.
- January 24: Presentation at the iDASH Workshop about the secondary use of clinical images for biomedical research.
2010
- December: Survey of IGT activities
- November 8-9: West Coast Events
- November 5-6: Algorithm Core retreat
- November 4-6: Widget Design Fiesta, Kitware, South Carborro, NC.
- September 28: Visit at UCLA: preparation for DBP activities, talk
- September 23: Slicer Tutorial at Chinese PLA General Hospital Beijing, China.
- September 20: NA-MIC Tutorial at MICCAI 2010, Beijing, China.
- August 26: Planning renewal activities
- August 5: Presentation at the AKH, Vienna, Austria.
- June 28: Presentation at the ITK 10th Anniversary.
- April 9-10: Slicer Training at the University of Iowa, Iowa City, IA.
- March 16: Presentation at the Slicer training events: Tokyo Women's Medical University, Tokyo, Japan.
- March 15: Presentations at Tokyo University, Department of Radiology, Department of Mechanical Engineering and Biomedical Engineering. See here for the presentation.
- March 11: Presentation at the AMIA translational bioinformatics workshop in SF. See here for the presentation.
- March 11: Visit and Slicer presentation at the SF, VA.
- Feb 9-11: Collaboration visit in SLC, Utah. Meeting with the autism and cardiac projects people.
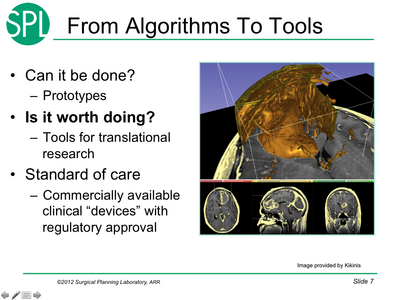
- February 5: Rons rules for tools
- January 8: NA-MIC competitive renewal submitted
2009
- October 1: Lecture at MIT biomedical computing class
- October 1: ARRA Supplements
- September 24: Ron Kikinis awarded the inaugural Enduring Impact Award of the MICCAI Society during the annual meeting, London, UK.
- September 10: Open Source Software as an Enabler of Science. Word Congress 2009 Medical Physics and Biomedical Engineering, Munich, Germany.
- June 29: Presentation about 3D Slicer. Working meeting about a common toolkit.
- June 18: Open Source Software as Enabler of Research. Presidential Guest Lecture at the 9th Annual Meeting of the International Society for Computer Assisted Orthopedic Surgery
- June 4: Presentation at MEVIS
- May 11-12: Core 1 visit to Georgia Tech: brainstorming for competitive renewal
- May 5-4: Core 2 visit to Georgia Tech: brainstorming for competitive renewal
- February 27: Visit at NCI to explore interactions with CaBIG and the intramural animal imaging program
2008
- October November December 2008: Multiple trips and meetings to scope out next generation DBP's
- September: MICCAI, Core 1 PI get together
- August: NCBC AHM meeting in Bethesda
- July: Planning of next generation DBPs begins
- June: Participation in the Project week
- June: Participation in the Germany outreach event
- April: Na-mic presentation at McMaster University
- March 13: Visit at Allen Brain Institute, Seattle, VA.
- March 12: Visit at Intuitive in Sunnyvale
- March 09: Presentation for the AMIA Translational Bioinformatics Summit
- January 17: Visit of Georgia Tech
- January 8: Slicer3 compilation discussion
2007
- October 25: Presentation about Open Source in Robotics
- October 12: Discussion about Slicer and IGT
- October 11: The Application of Open Source Concepts to Image Guided Therapy Plenary Session talk at CURAC
- October 3: Ron Kikinis presenting at IIC, Harvard: Medical Image Computing: From Data to Understanding, Video of the presentation, PPT of the presentation
- September 1-7: Publications database introduced
- September: Projects for the DBP's in 2007
- April 15: ISBI Plenary Session Lecture about MIC.
2006
- November 21: NA-MIC and NAC presentations at the DKFZ, Heidelberg, Germany.
- October 26: Visiting the MIND institute
- September 6: 2006 IBMISPS conference: talk by R. Kikinis about FOSS
- May 5: Meeting with Andy Cedilnik at 1249 Boylston to review plans for wiki based maintenance for the NA-MIC website
- April 24: The Human Brain Project: Linking Within and Beyond Natcher Conference Center, NIH Campus, Bethesda, Maryland: National Alliance for Medical Image Computing: Science & Technology of One National Center for Biomedical Computing (this is a slightly modified version of the presentation)
- April 18: Scientific Visualization 2006 Workshop, University of Minnesota, Minneapolis: 3D SLICER: An Open Source Tool for Subject-Specific Data Analysis abridged presentation ppt (20MB) snapshot of the three presenters
- April 8: ISBI conference SA-AM-SS1: Towards Vertical Integration of Biomedical Data presentation ppt
- March 23: TCON among Core PIs on the selection of next generation DBPs.
- March 17: TCON among Core PIs on the new format of annual scientific report by projects, not by cores.
- March 11-30: Pre-selection of next generation of NAMIC-DBPs and the phase out plan of existing ones
- January 8-13: The second All-Hands-Meeting and Programming event, Salt Lake City, UT.
2005
- December 12: Start NCBC Evaluation TCON with a total of seven NCBC centers
- November 18: Initial T-con with EAB
- September-October: Closing of accounts and preparing Carryforward Request
- July: External Advisory Board (EAB) is formed
- May-June: Coordination with all sites to prepare the Annual Progress Report as well as Carryforward explanations.
- May-Oct: Interaction with ISC (ITK Software Consortium) as well as Partners OGC (Office of General Counsel) about Slicer Licensing
- April: NAMIC management "office hours"
- April 9: Coordination of Core 3, prospective IRB issues (CAMH, UCI, and Dartmouth no need for new IRBs. VA/Harvard filing soon)
- March 30: NAMIC accomplishment snapshot for NIH
- Feb 1-18: Harvard, Dartmouth and UCI data cleaned, validated and uploaded
- February 7: Attended NIH-NCBC Evaluation Liaisons Conference Call Meeting
- January 12: Secondary-use IRB approved by Partners
2004
- December 21: Filed Secondary-use IRB for data-sharing with Partners' IRB (for use of VA/Harvard and Dartmouth data)
- December 17: Dartmouth's Secondary-use IRB for restrospective data approved for sharing with NAMIC
- December 10: Coordinated with MIT/VA/Harvard to start a straw-man proposal for data-sharing policy for NAMIC
- December 8: Attend NCBC meeting in NIH
- November 30: VA/Harvard's Secondary-use IRB for retrospective data approved for sharing with NAMIC
- November 20: Finalized schedule for all-hand meeting on Feb 5 2004, Salt Lake City, Utah
- November 7: Defined master project plan for NAMIC Year 1 activities and worked with Core 4 to post it on NAMIC website.
- November 3: Identified two potential collaborative projects, one in diGeorge brain MRI imaging genome and the other in COPD lung cancer CT imaging genome with investigators from I2B2, another NCBC at Harvard.
- October 15: Host first NAMIC kickoff meeting @ Stata Center, MIT, MA
- October 14: Meet with NIH program managers and Art Toga--the PI of another NCBC imaging related Center, BWH Radiology Boylston Laboratory
- October 10: Start working with all cores on the multi-institutional IRBs for NAMIC.
- October 5: Leadership core planning on NAMIC kick-off meeting
- October 1: Set up NAMIC emailing groups